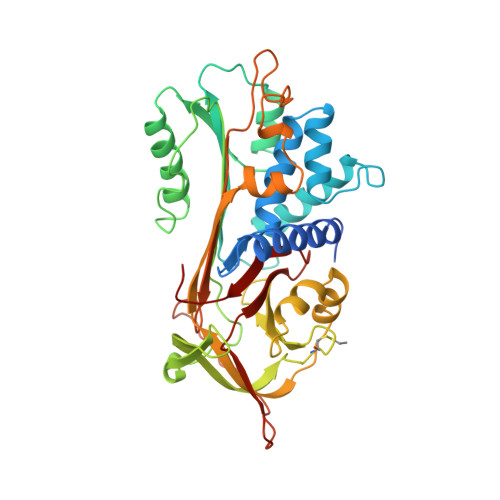
Inhibitory conformation of the reactive loop of alpha 1-antitrypsin.
Elliott, P.R., Lomas, D.A., Carrell, R.W., Abrahams, J.P.(1996) Nat Struct Biol 3: 676-681
- PubMed: 8756325
- DOI: https://doi.org/10.1038/nsb0896-676
- Primary Citation of Related Structures:
1PSI - PubMed Abstract:
The reactive site loop of the serpin family of serine proteinase inhibitors is flexible and can adopt a number of diverse conformations. A 2.9 A resolution structure of alpha 1-antitrypsin-the principal proteinase inhibitor in human plasma-shows the loop in a stable canonical conformation matching that found in all other families of serine proteinase inhibitors. This unexpected finding in the absence of loop insertion into the body of the molecule favours a two-stage mechanism of inhibition and provides a model for the heparin activation of antithrombin. The beta-pleated strand conformation of the loop also accounts for the polymerization of the serpins in disease and for their association with other beta-sheet structures, most notably the beta-amyloid of Alzheimer's disease.
Organizational Affiliation:
Department of Haematology, University of Cambridge, UK.