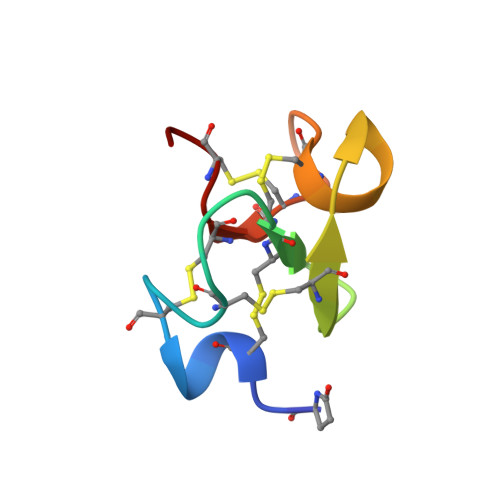
Solution Structure of Eucommia Antifungal Peptide: A Novel Structural Model Distinct with a Five-Disulfide Motif.
Huang, R.H., Xiang, Y., Tu, G.Z., Zhang, Y., Wang, D.C.(2004) Biochemistry 43: 6005-6012
- PubMed: 15147184
- DOI: https://doi.org/10.1021/bi036263y
- Primary Citation of Related Structures:
1P9Z - PubMed Abstract:
The three-dimensional structure in aqueous solution of Eucommia antifungal peptide 2 (EAFP2) from Eucommia ulmoides Oliv was determined using (1)H NMR spectroscopy. EAFP2 is a newly discovered 41-residue peptide distinct with a five-disulfide cross-linked motif. This peptide exhibits chitin-binding activity and inhibitory effects on the growth of cell wall chitin-containing fungi and chitin-free fungi. The structure was calculated by using torsion angle dynamic simulated annealing with a total of 614 distance restraints and 16 dihedral restraints derived from NOESY and DQF-COSY spectra, respectively. The five disulfide bonds were assigned from preliminary structures using a statistical analysis of intercystinyl distances. The solution structure of EAFP2 is presented as an ensemble of 20 conformers with a backbone RMS deviation of 0.65 (+/-0.13) A for the well-defined Cys3-Cys39 segment. The tertiary structure of EAFP2 represents the first five-disulfide cross-linked structural model of the plant antifungal peptide. EAFP2 adopts a compact global fold composed of a 3(10) helix (Cys3-Arg6), an alpha-helix (Gly26-Cys30), and a three-strand antiparallel beta-sheet (Cys16-Ser18, Tyr22-Gly24, and Arg36-Cys37). The tertiary structure of EAFP2 shows a chitin-binding domain (residues 11-30) with a hydrophobic face and a characteristic sector formed by the N-terminal 10 residues and the C-terminal segment cross-linked through the unique disulfide bond Cys7-Cys37, which brings all four positively charged residues (Arg6, Arg9, Arg36, and Arg40) onto a cationic face. On the basis of such a structural feature, the possible structural basis for the functional properties of EAFP2 is discussed.
Organizational Affiliation:
Center for Structural and Molecular Biology, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, PR China.