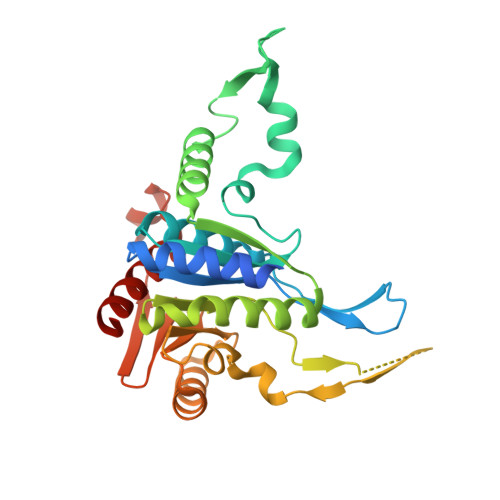
Structure of human phosphopantothenoylcysteine synthetase at 2.3 A resolution.
Manoj, N., Strauss, E., Begley, T.P., Ealick, S.E.(2003) Structure 11: 927-936
- PubMed: 12906824
- DOI: https://doi.org/10.1016/s0969-2126(03)00146-1
- Primary Citation of Related Structures:
1P9O - PubMed Abstract:
The structure of human phosphopantothenoylcysteine (PPC) synthetase was determined at 2.3 A resolution. PPC synthetase is a dimer with identical monomers. Some features of the monomer fold resemble a group of NAD-dependent enzymes, while other features resemble the ribokinase fold. The ATP, phosphopantothenate, and cysteine binding sites were deduced from modeling studies. Highly conserved ATP binding residues include Gly43, Ser61, Gly63, Gly66, Phe230, and Asn258. Highly conserved phosphopantothenate binding residues include Asn59, Ala179, Ala180, and Asp183 from one monomer and Arg55' from the adjacent monomer. The structure predicts a ping pong mechanism with initial formation of an acyladenylate intermediate, followed by release of pyrophosphate and attack by cysteine to form the final products PPC and AMP.
Organizational Affiliation:
Department of Chemistry and Chemical Biology, Cornell University, Ithaca, NY 14853, USA.