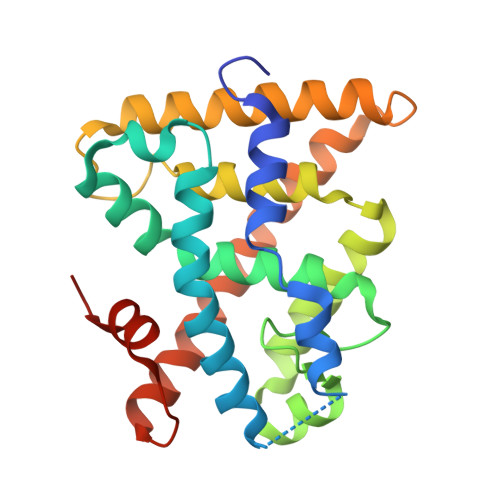
Molecular Recognition of Agonist Ligands by RXRs
Egea, P.F., Mitschler, A., Moras, D.(2002) Mol Endocrinol 16: 987-997
- PubMed: 11981034
- DOI: https://doi.org/10.1210/mend.16.5.0823
- Primary Citation of Related Structures:
1MV9, 1MVC, 1MZN - PubMed Abstract:
The nuclear receptor RXR is an obligate partner in many signal transduction pathways. We report the high-resolution structures of two complexes of the human RXRalpha ligand-binding domain specifically bound to two different and chemically unrelated agonist compounds: docosa hexaenoic acid, a natural derivative of eicosanoic acid, present in mammalian cells and recently identified as a potential endogenous RXR ligand in the mouse brain, and the synthetic ligand BMS 649. In both structures the RXR-ligand-binding domain forms homodimers and exhibits the active conformation previously observed with 9-cis-RA. Analysis of the differences in ligand-protein contacts (predominantly van der Waals forces) and binding cavity geometries and volumes for the several agonist-bound RXR structures clarifies the structural features important for ligand recognition. The L-shaped ligand-binding pocket adapts to the diverse ligands, especially at the level of residue N306, which might thus constitute a new target for drug-design. Despite its highest affinity 9-cis-RA displays the lowest number of ligand-protein contacts. These structural results support the idea that docosa hexaenoic acid and related fatty acids could be natural agonists of RXRs and question the real nature of the endogenous ligand(s) in mammalian cells.
Organizational Affiliation:
Laboratoire de Biologie et Génomique Structurales, Université Louis Pasteur, Parc d'Innovation BP163, 67404 Illkirch cedex, France.