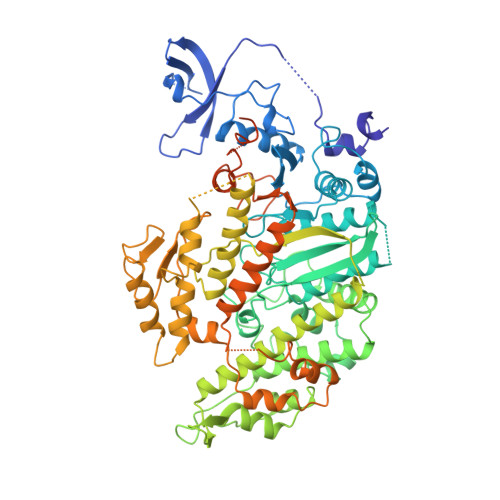
X-ray structures of the myosin motor domain of Dictyostelium discoideum complexed with MgADP.BeFx and MgADP.AlF4-.
Fisher, A.J., Smith, C.A., Thoden, J.B., Smith, R., Sutoh, K., Holden, H.M., Rayment, I.(1995) Biochemistry 34: 8960-8972
- PubMed: 7619795
- DOI: https://doi.org/10.1021/bi00028a004
- Primary Citation of Related Structures:
1MMD, 1MND - PubMed Abstract:
The three-dimensional structures of the truncated myosin head from Dictyostelium discoideum myosin II complexed with beryllium and aluminum fluoride and magnesium ADP are reported at 2.0 and 2.6 A resolution, respectively. Crystals of the beryllium fluoride-MgADP complex belong to space group P2(1)2(1)2 with unit cell parameters of a = 105.3 A, b = 182.6 A, and c = 54.7 A, whereas the crystals of the aluminum fluoride complex belong to the orthorhombic space group C222(1) with unit cell dimensions of a = 87.9 A, b = 149.0 A, and c = 153.8 A. Chemical modification was not necessary to obtain these crystals. These structures reveal the location of the nucleotide complexes and define the amino acid residues that form the active site. The tertiary structure of the protein complexed with MgADP.BeFx is essentially identical to that observed previously in the three-dimensional model of chicken skeletal muscle myosin subfragment-1 in which no nucleotide was present. By contrast, the complex with MgADP.AlF4- exhibits significant domain movements. The structures suggest that the MgADP.BeFx complex mimics the ATP bound state and the MgADP.AlF4- complex is an analog of the transition state for hydrolysis. The domain movements observed in the MgADP.AlF4- complex indicate that myosin undergoes a conformational change during hydrolysis that is not associated with the nucleotide binding pocket but rather occurs in the COOH-terminal segment of the myosin motor domain.
Organizational Affiliation:
Institute for Enzyme Research, Department of Biochemistry, University of Wisconsin, Madison 53705, USA.