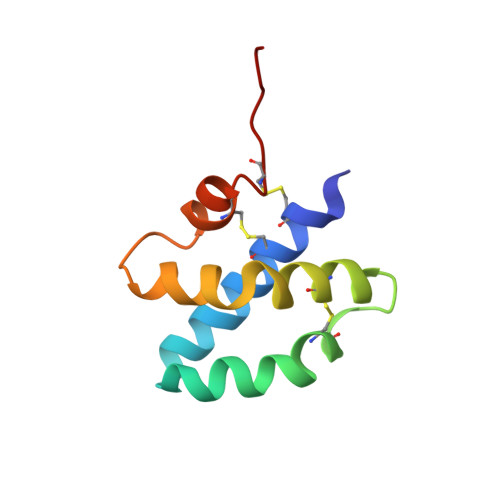
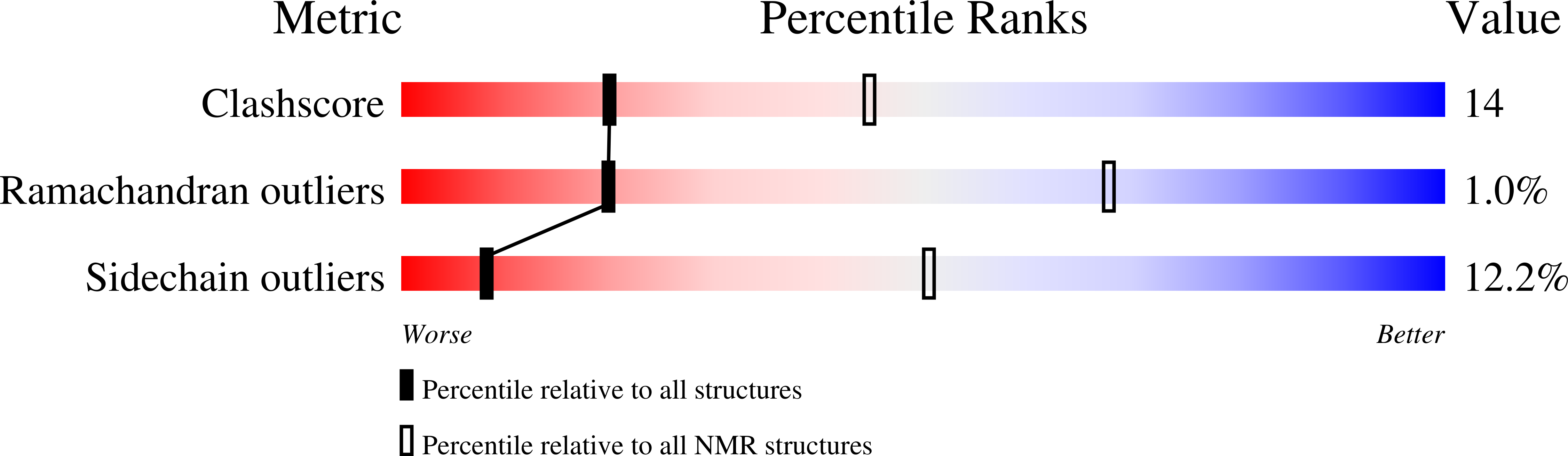Solution structure of human saposin C: pH-dependent interaction with phospholipid vesicles.
de Alba, E., Weiler, S., Tjandra, N.(2003) Biochemistry 42: 14729-14740
- PubMed: 14674747
- DOI: https://doi.org/10.1021/bi0301338
- Primary Citation of Related Structures:
1M12 - PubMed Abstract:
Saposin C binds to membranes to activate lipid degradation in lysosomes. To get insights into saposin C's function, we have determined its three-dimensional structure by NMR and investigated its interaction with phospholipid vesicles. Saposin C adopts the saposin-fold common to other members of the family. In contrast, the electrostatic surface revealed by the NMR structure is remarkably different. We suggest that charge distribution in the protein surface can modulate membrane interaction leading to the functional diversity of this family. We find that the binding of saposin C to phospholipid vesicles is a pH-controlled reversible process. The pH dependence of this interaction is sigmoidal, with an apparent pK(a) for binding close to 5.3. The pK(a) values of many solvent-exposed Glu residues are anomalously high and close to the binding pK(a). Our NMR data are consistent with the absence of a conformational change prior to membrane binding. All this information suggests that the negatively charged electrostatic surface of saposin C needs to be partially neutralized to trigger membrane binding. We have studied the membrane-binding behavior of a mutant of saposin C designed to decrease the negative charge of the electrostatic surface. The results support our conclusion on the importance of protein surface neutralization in binding. Since saposin C is a lysosomal protein and pH gradients occur in lysosomes, we propose that lipid degradation in the lysosome could be switched on and off by saposin C's reversible binding to membranes.
Organizational Affiliation:
Laboratory of Biophysical Chemistry, National Heart, Lung, and Blood Institute, National Institutes of Health, 50 Center Drive, Bethesda, Maryland 20892, USA. dealba@helix.nih.gov