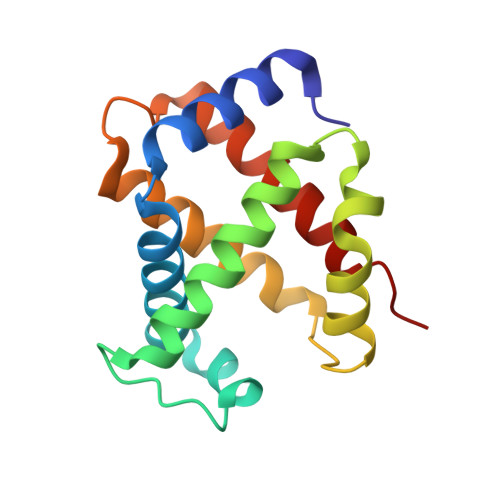
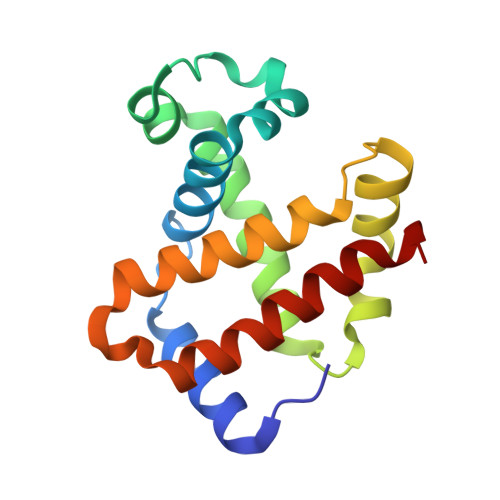
Structure of human carbonmonoxyhemoglobin at 2.16 A: a snapshot of the allosteric transition.
Safo, M.K., Burnett, J.C., Musayev, F.N., Nokuri, S., Abraham, D.J.(2002) Acta Crystallogr D Biol Crystallogr 58: 2031-2037
- PubMed: 12454461
- DOI: https://doi.org/10.1107/s0907444902015809
- Primary Citation of Related Structures:
1LJW - PubMed Abstract:
A 2.16 A resolution structure of high-salt human carbonmonoxyhemoglobin crystallized at pH 6.4 is reported. The quaternary structure is similar to that of 'classic' R-state hemoglobin; however, subtle but significant tertiary structural changes are observed at the alpha(1)beta(2) and symmetrically equivalent alpha(2)beta(1) interfaces--these are the key subunit interfaces that govern the allosteric transition between the R and T states. Specifically, the movement and weakening of two important hydrogen bonds that are diagnostic for R-state structures, beta(2)His97-alpha(1)Thr38 and beta(2)Arg40-alpha(1)Thr41, have been observed. In addition, a phosphate molecule bound between the two beta-subunits (at the entrance to the central water cavity) has been identified and electron density indicates that this molecule occupies two alternate positions that are related by the dyad axis. Both positions superimpose on the 2,3-diphosphoglycerate binding site. One phosphate conformer interacts with beta(2)Asn139, beta(1)His143 and beta(1)His146, while the second interacts with symmetry-related counterparts (beta(1)Asn139, beta(2)His143 and beta(2)His146).
Organizational Affiliation:
School of Pharmacy, Department of Medicinal Chemistry and Institute for Structural Biology and Drug Discovery, Virginia Commonwealth University, Richmond, VA 23219, USA. msafo@hsc.vcu.edu