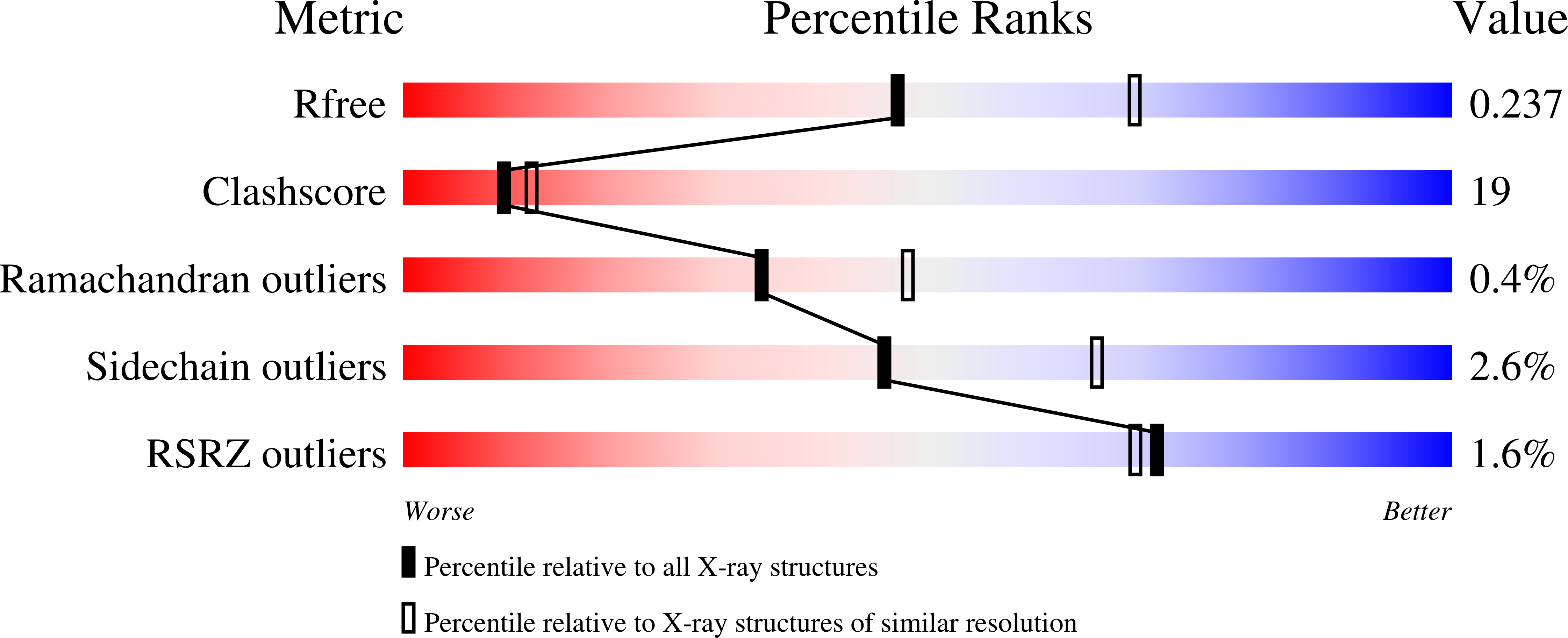
The Crystal Structure of the Allosteric Non-phosphorylating glyceraldehyde-3-phosphate Dehydrogenase from the Hyperthermophilic Archaeum Thermoproteus tenax
Pohl, E., Brunner, N., Wilmanns, M., Hensel, R.(2002) J Biol Chem 277: 19938-19945
- PubMed: 11842090
- DOI: https://doi.org/10.1074/jbc.M112244200
- Primary Citation of Related Structures:
1KY8 - PubMed Abstract:
The NAD(+)-dependent non-phosphorylating glyceraldehyde-3-phosphate dehydrogenase (GAPN) from the hyperthermophilic archaeum Thermoproteus tenax represents an archaeal member of the diverse superfamily of aldehyde dehydrogenases (ALDHs). GAPN catalyzes the irreversible oxidation of d-glyceraldehyde 3-phosphate to 3-phosphoglycerate. In this study, we present the crystal structure of GAPN in complex with its natural inhibitor NADP(+) determined by multiple anomalous diffraction methods. The structure was refined to a resolution of 2.4 A with an R-factor of 0.21. The overall fold of GAPN is similar to the structures of ALDHs described previously, consisting of three domains: a nucleotide-binding domain, a catalytic domain, and an oligomerization domain. Local differences in the active site are responsible for substrate specificity. The inhibitor NADP(+) binds at an equivalent site to the cosubstrate-binding site of other ALDHs and blocks the enzyme in its inactive state, possibly preventing the transition to the active conformation. Structural comparison between GAPN from the hyperthermophilic T. tenax and homologs of mesophilic organisms establishes several characteristics of thermostabilization. These include protection against heat-induced covalent modifications by reducing and stabilizing labile residues, a decrease in number and volume of empty cavities, an increase in beta-strand content, and a strengthening of subunit contacts by ionic and hydrophobic interactions.
Organizational Affiliation:
European Molecular Biology Laboratory, Hamburg Outstation, Notkestrasse 85, D-22603 Hamburg, Germany. ehmke@embl-hamburg.de