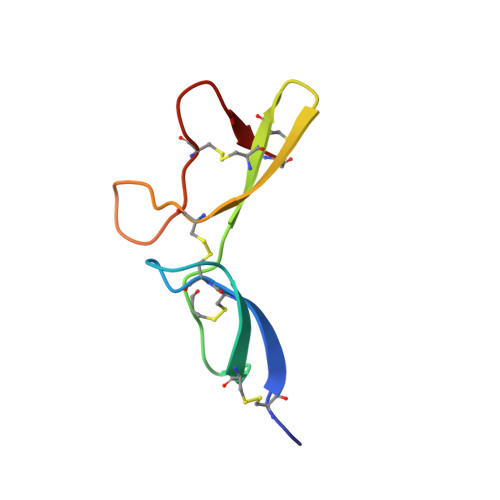
Solution Structure of the Plant Disease Resistance-triggering Protein NIP1 from the Fungus Rhynchosporium secalis Shows a Novel beta-Sheet Fold.
Van't Slot, K.A., Van Den Burg, H.A., Kloks, C.P., Hilbers, C.W., Knogge, W., Papavoine, C.H.(2003) J Biol Chem 278: 45730-45736
- PubMed: 12944393
- DOI: https://doi.org/10.1074/jbc.M308304200
- Primary Citation of Related Structures:
1KG1 - PubMed Abstract:
Activation of the disease resistance response in a host plant frequently requires the interaction of a plant resistance gene product with a corresponding, pathogenderived signal encoded by an avirulence gene. The products of resistance genes from diverse plant species show remarkable structural similarity. However, due to the general paucity of information on pathogen avirulence genes the recognition process remains in most cases poorly understood. NIP1, a small protein secreted by the fungal barley pathogen Rhynchosporium secalis, is one of only a few fungal avirulence proteins identified and characterized to date. The defense-activating activity of NIP1 is mediated by barley resistance gene Rrs1. In addition, a role of the protein in fungal virulence is suggested by its nonspecific toxicity in leaf tissues of host and non-host cereals as well as its resistance gene-independent stimulatory effect on the plant plasma membrane H+-ATPase. Four naturally occurring NIP1 isoforms are characterized by single amino acid alterations that affect the different activities in a similar way. As a step toward unraveling the signal perception/transduction mechanism, the solution structure of NIP1 was determined. The protein structure is characterized by a novel fold. It consists of two parts containing beta-sheets of two and three anti-parallel strands, respectively. Five intramolecular disulfide bonds, comprising a novel disulfide bond pattern, stabilize these parts and their position with respect to each other. A comparative analysis of the protein structure with the properties of the NIP1 isoforms suggests two loop regions to be crucial for the resistance-triggering activity of NIP1.
Organizational Affiliation:
Laboratory of Phytopathology, Department of Plant Sciences, Wageningen University, Binnenhaven 5, NL-6709 PD Wageningen, The Netherlands.