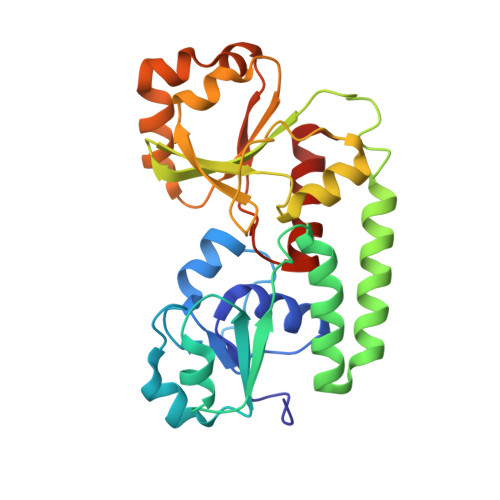
X-ray crystallographic structures of the Escherichia coli periplasmic protein FhuD bound to hydroxamate-type siderophores and the antibiotic albomycin.
Clarke, T.E., Braun, V., Winkelmann, G., Tari, L.W., Vogel, H.J.(2002) J Biol Chem 277: 13966-13972
- PubMed: 11805094
- DOI: https://doi.org/10.1074/jbc.M109385200
- Primary Citation of Related Structures:
1ESZ, 1K2V, 1K7S - PubMed Abstract:
Siderophore-binding proteins play an essential role in the uptake of iron in many Gram-positive and Gram-negative bacteria. FhuD is an ATP-binding cassette-type (ABC-type) binding protein involved in the uptake of hydroxamate-type siderophores in Escherichia coli. Structures of FhuD complexed with the antibiotic albomycin, the fungal siderophore coprogen and the drug Desferal have been determined at high resolution by x-ray crystallography. FhuD has an unusual bilobal structure for a periplasmic ligand binding protein, with two mixed beta/alpha domains connected by a long alpha-helix. The binding site for hydroxamate-type ligands is composed of a shallow pocket that lies between these two domains. Recognition of siderophores primarily occurs through interactions between the iron-hydroxamate centers of each siderophore and the side chains of several key residues in the binding pocket. Rearrangements of side chains within the binding pocket accommodate the unique structural features of each siderophore. The backbones of the siderophores are not involved in any direct interactions with the protein, demonstrating how siderophores with considerable chemical and structural diversity can be bound by FhuD. For albomycin, which consists of an antibiotic group attached to a hydroxamate siderophore, electron density for the antibiotic portion was not observed. Therefore, this study provides a basis for the rational design of novel bacteriostatic agents, in the form of siderophore-antibiotic conjugates that can act as "Trojan horses," using the hydroxamate-type siderophore uptake system to actively deliver antibiotics directly into targeted pathogens.
Organizational Affiliation:
Structural Biology Research Group, Department of Biological Sciences, University of Calgary, 2500 University Dr. N.W., Calgary, Alberta T2N 1N4, Canada.