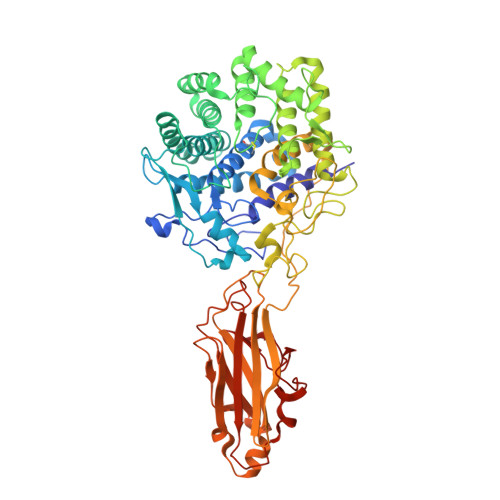
X-Ray Crystal Structure of the Multidomain Endoglucanase Cel9G from Clostridium cellulolyticum Complexed with Natural and Synthetic Cello-Oligosaccharides
Mandelman, D., Belaich, A., Belaich, J.P., Aghajari, N., Driguez, H., Haser, R.(2003) J Bacteriol 185: 4127-4135
- PubMed: 12837787
- DOI: https://doi.org/10.1128/JB.185.14.4127-4135.2003
- Primary Citation of Related Structures:
1G87, 1GA2, 1K72, 1KFG - PubMed Abstract:
Complete cellulose degradation is the first step in the use of biomass as a source of renewable energy. To this end, the engineering of novel cellulase activity, the activity responsible for the hydrolysis of the beta-1,4-glycosidic bonds in cellulose, is a topic of great interest. The high-resolution X-ray crystal structure of a multidomain endoglucanase from Clostridium cellulolyticum has been determined at a 1.6-A resolution. The endoglucanase, Cel9G, is comprised of a family 9 catalytic domain attached to a family III(c) cellulose-binding domain. The two domains together form a flat platform onto which crystalline cellulose is suggested to bind and be fed into the active-site cleft for endolytic hydrolysis. To further dissect the structural basis of cellulose binding and hydrolysis, the structures of Cel9G in the presence of cellobiose, cellotriose, and a DP-10 thio-oligosaccharide inhibitor were resolved at resolutions of 1.7, 1.8, and 1.9 A, respectively.
Organizational Affiliation:
Laboratoire de BioCristallographie, Institut de Biologie et Chimie des Protéines, UMR 5086, CNRS et Université Claude Bernard Lyon I, 69367 Lyon, Cedex 07, France.