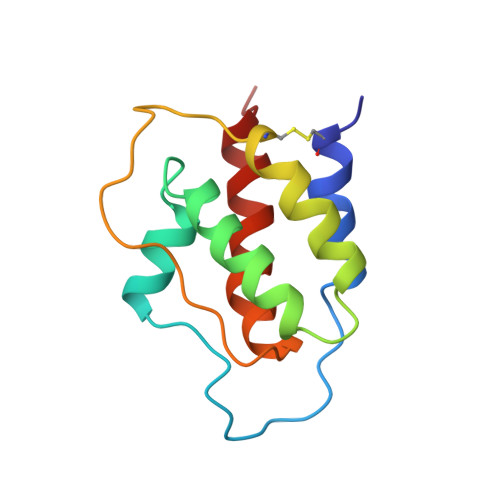
Three-dimensional solution structure and backbone dynamics of a variant of human interleukin-3.
Feng, Y., Klein, B.K., McWherter, C.A.(1996) J Mol Biol 259: 524-541
- PubMed: 8676386
- DOI: https://doi.org/10.1006/jmbi.1996.0337
- Primary Citation of Related Structures:
1JLI - PubMed Abstract:
The three-dimensional structure and backbone dynamics of a truncated and multiply substituted recombinant human interleukin-3 (IL-3) variant (SC-65369) have been determined from multidimensional heteronuclear nuclear magnetic resonance spectroscopic data. Sequential application of distance geometry and restrained molecular dynamics calculations produced a family of 25 convergent structures which satisfy a total of 1812 experimental constraints (1659 proton-proton NOEs, 75 backbone dihedral angle constraints, and 39 pairs of hydrogen bond constraints) with an average root-mean-square deviation from the mean coordinate positions of 0.88(+/- 0.15) angstroms and 1.37(+/- 0.13) angstroms for the backbone and all heavy atoms, respectively, of all residues except 28 to 39. The structure is a left-handed four-helix bundle (comprised of helices A through D) with two long overhand loops (designated as loops AB and CD). Loop AB contains a short fifth helix (helix A') which is closely packed against helix D in an approximately parallel fashion and which has multiple contacts with loop CD. The overall molecular tumbling time (6.5 ns) determined from the 15N relaxation data was consistent with a monomeric protein under the conditions of the experiment (1 mM protein, pH 4.6, 30 degrees C). The 15N relaxation data indicate that the helical regions of SC-65369 are quite rigid, while portions of loop AB, loop CD, and the C terminus undergo significant internal motions. Among the structurally related four-helical bundle cytokines, the structure of SC-65369 is most similar to those of granulocyte-macrophage colony stimulating factor (GM-CSF) and the single structural domain of interleukin-5 (IL-5), all of which share a common receptor subunit required for signal transduction and activation of their hematopoietic target cells. Indeed, the C(alpha) atoms in the four-helix core of these three proteins can be superimposed to 1.71 angstroms (SC-65369 and GM-CSF, 62 C(alpha) atoms) and 1.96 angstroms (SC-65369 and IL-5 single structural domain, 58 C(alpha) atoms), respectively. When the structures of the IL-3 variant, GM-CSF, and IL-5 were aligned, the conserved and conservatively substituted residues were found to be hydrophobic and buried, with the single exception of Glu-22 (IL-3 numbering), which is strictly conserved but nonetheless fully exposed to solvent. The most remarkable differences between the SC-65369 structure and that of GM-CSF occur in loop AB. This loop in GM-CSF crosses over the top of helix D and passes underneath loop CD on its way to helix B. In contrast, loop AB of SC-65369 passes in front of helix D, similar to the first crossover loop in human growth hormone and granulocyte colony-stimulating factor. In addition, helix A', which is interdigitated into the helical bundle in a manner similar to the helices in the CD loop of interferon-beta and interferon-gamma, exists in a region where short stretches of beta-structure are found at analogous positions in GM-CSF and IL-5. These differences suggest that the structural elements within this region may be important for recognition by their cognate receptors.
Organizational Affiliation:
G.D. Searle and Company, St. Louis, MO 63198, USA.