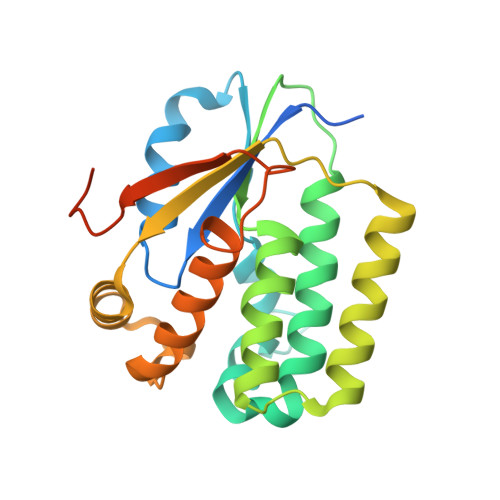
Structural basis for substrate specificities of cellular deoxyribonucleoside kinases.
Johansson, K., Ramaswamy, S., Ljungcrantz, C., Knecht, W., Piskur, J., Munch-Petersen, B., Eriksson, S., Eklund, H.(2001) Nat Struct Biol 8: 616-620
- PubMed: 11427893
- DOI: https://doi.org/10.1038/89661
- Primary Citation of Related Structures:
1J90, 2OCP - PubMed Abstract:
Deoxyribonucleoside kinases phosphorylate deoxyribonucleosides and activate a number of medically important nucleoside analogs. Here we report the structure of the Drosophila deoxyribonucleoside kinase with deoxycytidine bound at the nucleoside binding site and that of the human deoxyguanosine kinase with ATP at the nucleoside substrate binding site. Compared to the human kinase, the Drosophila kinase has a wider substrate cleft, which may be responsible for the broad substrate specificity of this enzyme. The human deoxyguanosine kinase is highly specific for purine substrates; this is apparently due to the presence of Arg 118, which provides favorable hydrogen bonding interactions with the substrate. The two new structures provide an explanation for the substrate specificity of cellular deoxyribonucleoside kinases.
Organizational Affiliation:
Department of Molecular Biology, Swedish University of Agricultural Sciences, Box 590, Biomedical Center, S-751 24 Uppsala, Sweden.