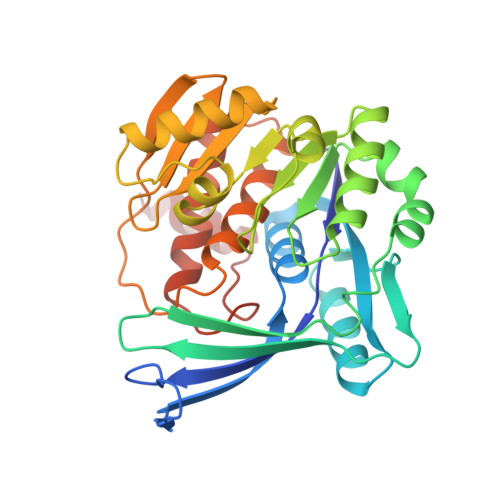
Activation of Ribokinase by Monovalent Cations.
Andersson, C.E., Mowbray, S.L.(2002) J Mol Biol 315: 409
- PubMed: 11786021
- DOI: https://doi.org/10.1006/jmbi.2001.5248
- Primary Citation of Related Structures:
1GQT - PubMed Abstract:
Carbohydrate kinases frequently require a monovalent cation for their activity. The physical basis of this phenomenon is, however, usually unclear. We report here that Escherichia coli ribokinase is activated by potassium with an apparent K(d) of 5 mM; the enzyme should therefore be fully activated under physiological conditions. Cesium can be used as an alternative ion, with an apparent K(d) of 17 mM. An X-ray structure of ribokinase in the presence of cesium was solved and refined at 2.34 A resolution. The cesium ion was bound between two loops immediately adjacent to the anion hole of the active site. The buried location of the site suggests that conformational changes will accompany ion binding, thus providing a direct mechanism for activation. Comparison with structures of a related enzyme, the adenosine kinase of Toxoplasma gondii, support this proposal. This is apparently the first instance in which conformational activation of a carbohydrate kinase by a monovalent cation has been assigned a clear structural basis. The mechanism is probably general to ribokinases, to some adenosine kinases, and to other members of the larger family. A careful re-evaluation of the biochemical and structural data is suggested for other enzyme systems.
Organizational Affiliation:
Department of Cell and Molecular Biology, Uppsala University, S-751 24 Uppsala, Sweden.