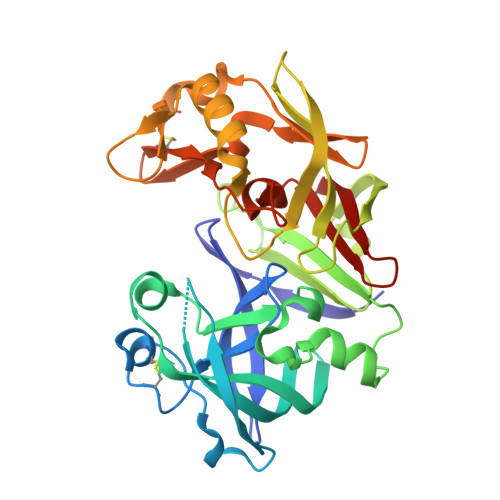
An unusual orientation for Tyr75 in the active site of the aspartic proteinase from Saccharomyces cerevisiae.
Gustchina, A., Li, M., Phylip, L.H., Lees, W.E., Kay, J., Wlodawer, A.(2002) Biochem Biophys Res Commun 295: 1020-1026
- PubMed: 12127998
- DOI: https://doi.org/10.1016/s0006-291x(02)00742-8
- Primary Citation of Related Structures:
1FMU, 1FMX - PubMed Abstract:
The structures of the native Saccharomyces cerevisiae proteinase A have been solved by molecular replacement in the monoclinic and trigonal crystal forms and refined at 2.6-2.7A resolution. These structures agree overall with those of other uninhibited aspartic proteinases. However, an unusual orientation for the side chain of Tyr75, a conserved residue on the flexible "flap" that covers the active site and is important for the activity of these enzymes, was found in the trigonal crystals. A similar conformation of Tyr75 occupying the S1 substrate-binding pocket was previously reported only for chymosin (where it was interpreted as representing a "self-inhibited" state of the enzyme), but for no other aspartic proteinases. Since this orientation of Tyr75 has now been seen in the structures of two members of the family of aspartic proteinases, it might indicate that the placement of that residue in the S1 substrate-binding pocket might have some functional significance, analogous to what was seen for self-inhibited structures of serine proteinases.
Organizational Affiliation:
Protein Structure Section, Macromolecular Crystallography Laboratory, National Cancer Institute at Frederick, Frederick, MD 21702-1201, USA.