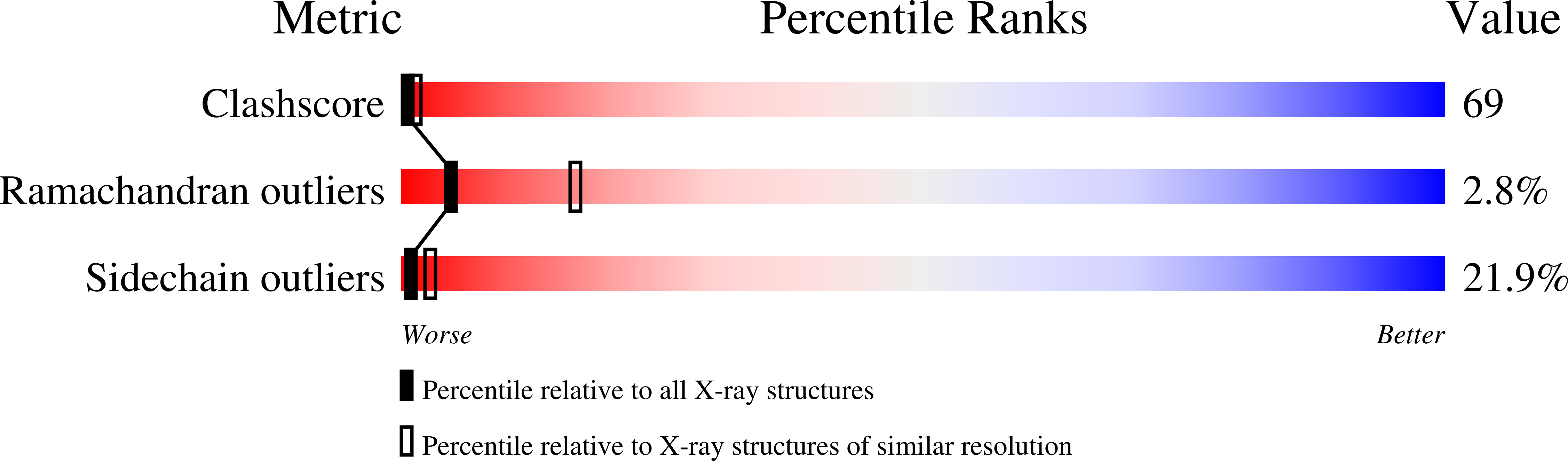Effects of site-directed mutagenesis on structure and function of recombinant rat liver S-adenosylhomocysteine hydrolase. Crystal structure of D244E mutant enzyme.
Komoto, J., Huang, Y., Gomi, T., Ogawa, H., Takata, Y., Fujioka, M., Takusagawa, F.(2000) J Biol Chem 275: 32147-32156
- PubMed: 10913437
- DOI: https://doi.org/10.1074/jbc.M003725200
- Primary Citation of Related Structures:
1D4F - PubMed Abstract:
A site-directed mutagenesis, D244E, of S-adenosylhomocysteine hydrolase (AdoHcyase) changes drastically the nature of the protein, especially the NAD(+) binding affinity. The mutant enzyme contained NADH rather than NAD(+) (Gomi, T., Takata, Y., Date, T., Fujioka, M., Aksamit, R. R., Backlund, P. S., and Cantoni, G. L. (1990) J. Biol. Chem. 265, 16102-16107). In contrast to the site-directed mutagenesis study, the crystal structures of human and rat AdoHcyase recently determined have shown that the carboxyl group of Asp-244 points in a direction opposite to the bound NAD molecule and does not participate in any hydrogen bonds with the NAD molecule. To explain the discrepancy between the mutagenesis study and the x-ray studies, we have determined the crystal structure of the recombinant rat-liver D244E mutant enzyme to 2.8-A resolution. The D244E mutation changes the enzyme structure from the open to the closed conformation by means of a approximately 17 degrees rotation of the individual catalytic domains around the molecular hinge sections. The D244E mutation shifts the catalytic reaction from a reversible to an irreversible fashion. The large affinity difference between NAD(+) and NADH is mainly due to the enzyme conformation, but not to the binding-site geometry; an NAD(+) in the open conformation is readily released from the enzyme, whereas an NADH in the closed conformation is trapped and cannot leave the enzyme. A catalytic mechanism of AdoHcyase has been proposed on the basis of the crystal structures of the wild-type and D244E enzymes.
Organizational Affiliation:
Department of Molecular Biosciences, University of Kansas, Lawrence, Kansas 66045-210, USA.