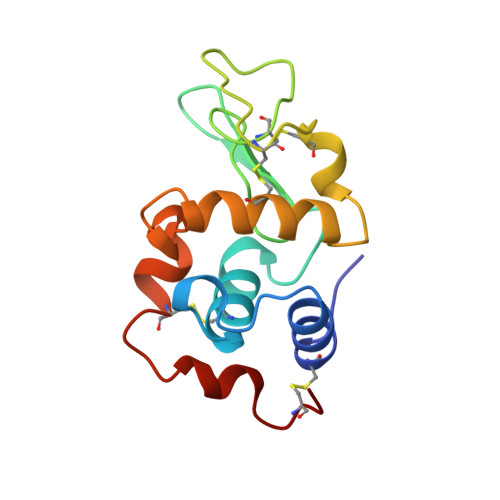
Refined structure of orthorhombic lysozyme crystallized at high temperature: correlation between morphology and intermolecular contacts.
Oki, H., Matsuura, Y., Komatsu, H., Chernov, A.A.(1999) Acta Crystallogr D Biol Crystallogr 55: 114-121
- PubMed: 10089401
- DOI: https://doi.org/10.1107/S0907444998008713
- Primary Citation of Related Structures:
1BGI - PubMed Abstract:
The structure of orthorhombic hen egg-white lysozyme (HEWL) crystallized at 310 K has been refined at 1.7 A resolution. Large displacements of the side-chain atoms with respect to the tetragonal structure were observed in many places, in contrast to small displacements of the main-chain atoms. A chloride-ion binding site was observed at an interface of two molecules, but at a different position to the binding site in the tetragonal form. The analysis of intermolecular contacts in the crystal has shown the presence of three independent intermolecular contacts which are called macrobonds A, B and C. Arginine side chains are frequently involved in these macrobonds, suggesting that the high frequency of this residue in HEWL may be a possible reason for the multiple polymorphs of this protein. The crystal forms were determined using a light-reflecting device on a four-circle diffractometer. Correlations between crystal forms and the three-dimensional macrobond networks were interpreted in terms of their components in various crystallographic planes, making use of approximate strengths of hydrogen-bond and van der Waals interatomic forces.
Organizational Affiliation:
Institute for Protein Research, Osaka University, Suita, Osaka 565, Japan.