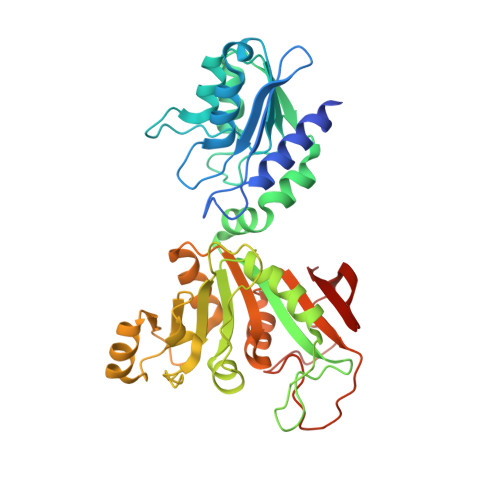
Complex structure of Bacillus subtilis RibG: the reduction mechanism during riboflavin biosynthesis.
Chen, S.C., Lin, Y.H., Yu, H.C., Liaw, S.H.(2009) J Biol Chem 284: 1725-1731
- PubMed: 18986985
- DOI: https://doi.org/10.1074/jbc.M805820200
- Primary Citation of Related Structures:
3EX8 - PubMed Abstract:
Bacterial RibG is a potent target for antimicrobial agents, because it catalyzes consecutive deamination and reduction steps in the riboflavin biosynthesis. In the N-terminal deaminase domain of Bacillus subtilis RibG, structure-based mutational analyses demonstrated that Glu51 and Lys79 are essential for the deaminase activity. In the C-terminal reductase domain, the complex structure with the substrate at 2.56-A resolution unexpectedly showed a ribitylimino intermediate bound at the active site, and hence suggested that the ribosyl reduction occurs through a Schiff base pathway. Lys151 seems to have evolved to ensure specific recognition of the deaminase product rather than the substrate. Glu290, instead of the previously proposed Asp199, would seem to assist in the proton transfer in the reduction reaction. A detailed comparison reveals that the reductase and the pharmaceutically important enzyme, dihydrofolate reductase involved in the riboflavin and folate biosyntheses, share strong conservation of the core structure, cofactor binding, catalytic mechanism, even the substrate binding architecture.
Organizational Affiliation:
Structural Biology Program, National Yang-Ming University, Taipei 11221, Taiwan.