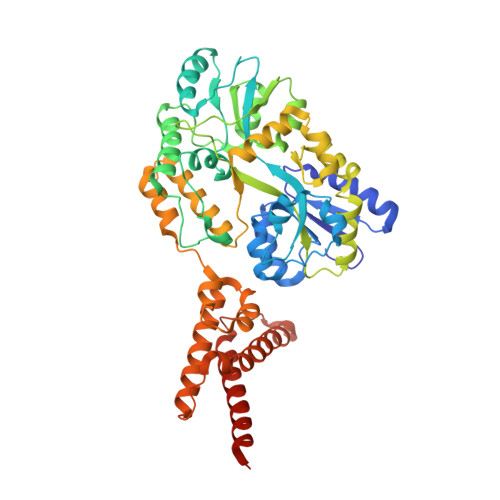
Mechanism of staphylococcal multiresistance plasmid replication origin assembly by the RepA protein.
Schumacher, M.A., Tonthat, N.K., Kwong, S.M., Chinnam, N.B., Liu, M.A., Skurray, R.A., Firth, N.(2014) Proc Natl Acad Sci U S A 111: 9121-9126
- PubMed: 24927575
- DOI: https://doi.org/10.1073/pnas.1406065111
- Primary Citation of Related Structures:
4PQK, 4PQL, 4PT7, 4PTA, 5KBJ - PubMed Abstract:
The staphylococcal multiresistance plasmids are key contributors to the alarming rise in bacterial multidrug resistance. A conserved replication initiator, RepA, encoded on these plasmids is essential for their propagation. RepA proteins consist of flexibly linked N-terminal (NTD) and C-terminal (CTD) domains. Despite their essential role in replication, the molecular basis for RepA function is unknown. Here we describe a complete structural and functional dissection of RepA proteins. Unexpectedly, both the RepA NTD and CTD show similarity to the corresponding domains of the bacterial primosome protein, DnaD. Although the RepA and DnaD NTD both contain winged helix-turn-helices, the DnaD NTD self-assembles into large scaffolds whereas the tetrameric RepA NTD binds DNA iterons using a newly described DNA binding mode. Strikingly, structural and atomic force microscopy data reveal that the NTD tetramer mediates DNA bridging, suggesting a molecular mechanism for origin handcuffing. Finally, data show that the RepA CTD interacts with the host DnaG primase, which binds the replicative helicase. Thus, these combined data reveal the molecular mechanism by which RepA mediates the specific replicon assembly of staphylococcal multiresistant plasmids.
Organizational Affiliation:
Department of Biochemistry, Duke University Medical Center, Durham, NC 27710; and maria.schumacher@duke.edu.