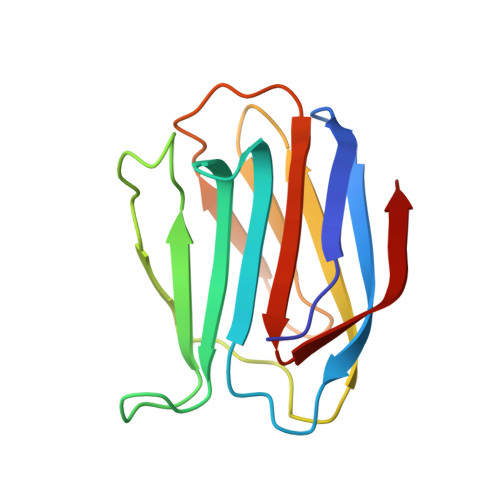
Lactose binding to galectin-1 modulates structural dynamics, increases conformational entropy, and occurs with apparent negative cooperativity.
Nesmelova, I.V., Ermakova, E., Daragan, V.A., Pang, M., Menendez, M., Lagartera, L., Solis, D., Baum, L.G., Mayo, K.H.(2010) J Mol Biol 397: 1209-1230
- PubMed: 20184898
- DOI: https://doi.org/10.1016/j.jmb.2010.02.033
- Primary Citation of Related Structures:
2KM2 - PubMed Abstract:
Galectins are a family of lectins with a conserved carbohydrate recognition domain that interacts with beta-galactosides. By binding cell surface glycoconjugates, galectin-1 (gal-1) is involved in cell adhesion and migration processes and is an important regulator of tumor angiogenesis. Here, we used heteronuclear NMR spectroscopy and molecular modeling to investigate lactose binding to gal-1 and to derive solution NMR structures of gal-1 in the lactose-bound and unbound states. Structure analysis shows that the beta-strands and loops around the lactose binding site, which are more open and dynamic in the unbound state, fold in around the bound lactose molecule, dampening internal motions at that site and increasing motions elsewhere throughout the protein to contribute entropically to the binding free energy. CD data support the view of an overall more open structure in the lactose-bound state. Analysis of heteronuclear single quantum coherence titration binding data indicates that lactose binds the two carbohydrate recognition domains of the gal-1 dimer with negative cooperativity, in that the first lactose molecule binds more strongly (K(1)=21+/-6 x 10(3) M(-1)) than the second (K(2)=4+/-2 x 10(3) M(-1)). Isothermal calorimetry data fit using a sequential binding model present a similar picture, yielding K(1)=20+/-10 x 10(3) M(-1) and K(2)=1.67+/-0.07 x 10(3) M(-1). Molecular dynamics simulations provide insight into structural dynamics of the half-loaded lactose state and, together with NMR data, suggest that lactose binding at one site transmits a signal through the beta-sandwich and loops to the second binding site. Overall, our results provide new insight into gal-1 structure-function relationships and to protein-carbohydrate interactions in general.
Organizational Affiliation:
Department of Biochemistry, Molecular Biology and Biophysics, University of Minnesota, Minneapolis, MN 55455, USA.