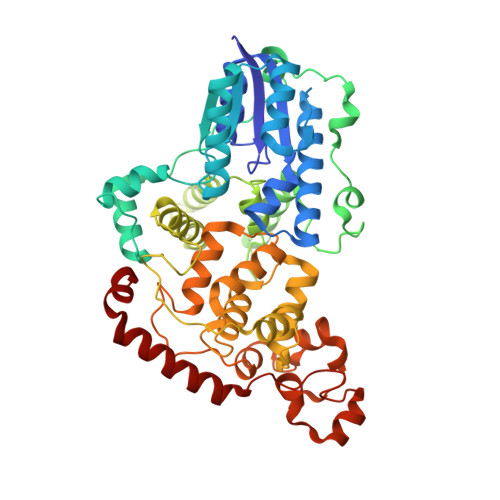
Chemical and structural analysis of a photoactive vertebrate cryptochrome from pigeon.
Zoltowski, B.D., Chelliah, Y., Wickramaratne, A., Jarocha, L., Karki, N., Xu, W., Mouritsen, H., Hore, P.J., Hibbs, R.E., Green, C.B., Takahashi, J.S.(2019) Proc Natl Acad Sci U S A 116: 19449-19457
- PubMed: 31484780 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1907875116
- Primary Citation Related Structures:
6PTZ, 6PU0 - PubMed Abstract:
Computational and biochemical studies implicate the blue-light sensor cryptochrome (CRY) as an endogenous light-dependent magnetosensor enabling migratory birds to navigate using the Earth's magnetic field. Validation of such a mechanism has been hampered by the absence of structures of vertebrate CRYs that have functional photochemistry. Here we present crystal structures of Columba livia (pigeon) CRY4 that reveal evolutionarily conserved modifications to a sequence of Trp residues (Trp-triad) required for CRY photoreduction. In Cl CRY4, the Trp-triad chain is extended to include a fourth Trp (W369) and a Tyr (Y319) residue at the protein surface that imparts an unusually high quantum yield of photoreduction. These results are consistent with observations of night migratory behavior in animals at low light levels and could have implications for photochemical pathways allowing magnetosensing.
- Department of Chemistry, Southern Methodist University, Dallas, TX 75275.
Organizational Affiliation: