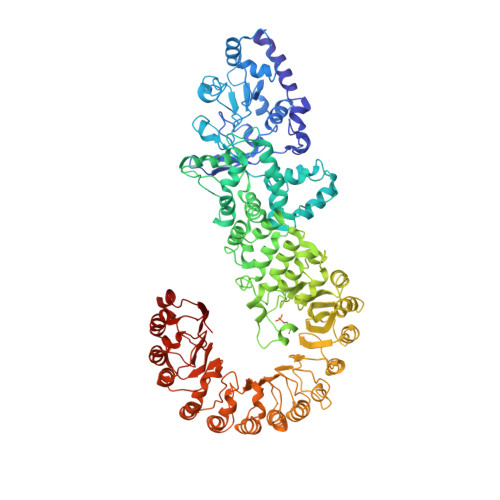
Cryo-EM structure of the activated NAIP2-NLRC4 inflammasome reveals nucleated polymerization.
Zhang, L., Chen, S., Ruan, J., Wu, J., Tong, A.B., Yin, Q., Li, Y., David, L., Lu, A., Wang, W.L., Marks, C., Ouyang, Q., Zhang, X., Mao, Y., Wu, H.(2015) Science 350: 404-409
- PubMed: 26449474 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.aac5789
- Primary Citation Related Structures:
3JBL - PubMed Abstract:
The NLR family apoptosis inhibitory proteins (NAIPs) bind conserved bacterial ligands, such as the bacterial rod protein PrgJ, and recruit NLR family CARD-containing protein 4 (NLRC4) as the inflammasome adapter to activate innate immunity. We found that the PrgJ-NAIP2-NLRC4 inflammasome is assembled into multisubunit disk-like structures through a unidirectional adenosine triphosphatase polymerization, primed with a single PrgJ-activated NAIP2 per disk. Cryo-electron microscopy (cryo-EM) reconstruction at subnanometer resolution revealed a ~90° hinge rotation accompanying NLRC4 activation. Unlike in the related heptameric Apaf-1 apoptosome, in which each subunit needs to be conformationally activated by its ligand before assembly, a single PrgJ-activated NAIP2 initiates NLRC4 polymerization in a domino-like reaction to promote the disk assembly. These insights reveal the mechanism of signal amplification in NAIP-NLRC4 inflammasomes.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115, USA. Program in Cellular and Molecular Medicine, Boston Children's Hospital, Boston, MA 02115, USA.
Organizational Affiliation: