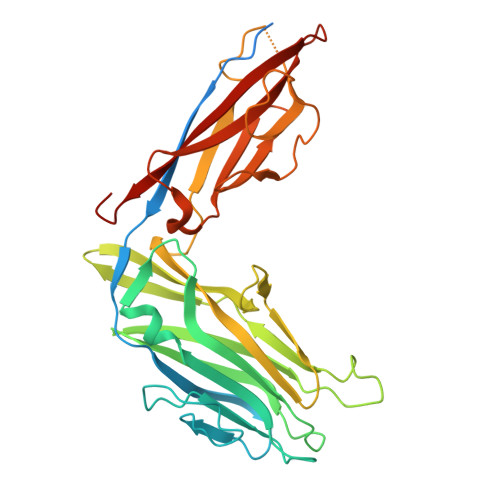
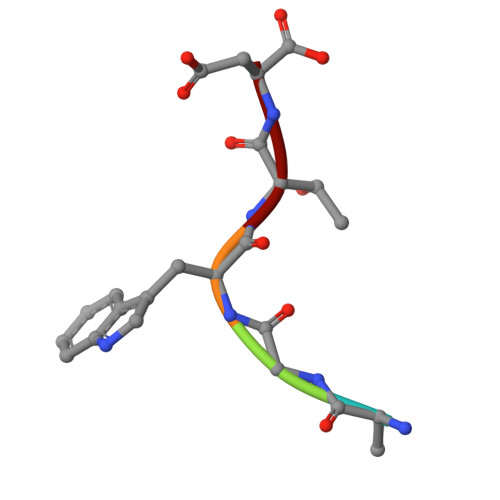
Peptide-based ligand antagonists block a Vibrio cholerae adhesin.
Wang, M., Du, G., Yongo-Luwawa, C., Lu, A., Kinrade, B., Munro, K., Klose, K.E., Lubell, W.D., Davies, P., Guo, S.(2025) FEBS Lett
- PubMed: 41262002
- DOI: https://doi.org/10.1002/1873-3468.70231
- Primary Citation of Related Structures:
9Y9W, 9YBQ - PubMed Abstract:
Vibrio cholerae, the causative agent of cholera, uses surface proteins such as the repeats-in-toxin (RTX) adhesin FrhA to colonize hosts and initiate infection. Blocking bacterial adhesion represents a promising therapeutic strategy to treat infections without promoting drug resistance. FrhA contains a peptide-binding domain (PBD) that is key for hemagglutination, human epithelial cell binding, and V. cholerae biofilm formation. Previous studies identified a lead pentapeptide ligand with the sequence Ala-Gly-Tyr-Thr-Asp (AGYTD) that blocks V. cholerae colonization of the mouse small intestine at high micromolar concentrations. In this study, a structure-guided approach identified a minimal D-amino acid-containing tripeptide motif with higher affinity for the FrhA-PBD and predicted metabolic stability. Our results contribute to the development of anti-adhesion strategies to combat infections. Impact statement Our study elucidates the molecular basis of peptide recognition by the Vibrio cholerae adhesin FrhA and develops minimal D-amino-acid peptides that block adhesion with nanomolar affinity. These findings advance understanding of RTX adhesins and provide a structural blueprint for next-generation anti-adhesion therapeutics against cholera and related infections.
- Department of Anatomy and Cell Biology, McGill University, Montreal, Canada.
Organizational Affiliation: