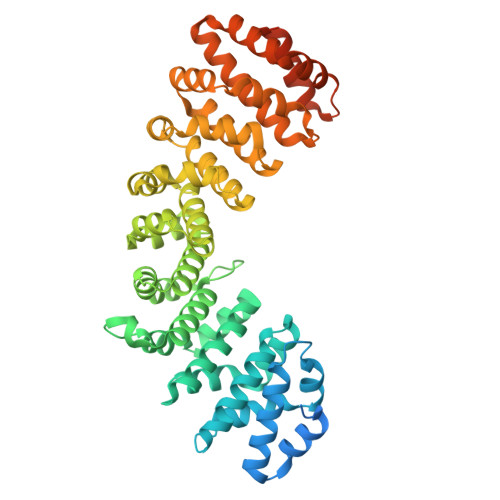
Structural and functional characterisation of ATF2 nuclear import reveals paralogue-selective importin-alpha recognition and a non-canonical NLS formed in trans.
Ghafoori, S.M., Pavan, S., Duc, T.X., Nematollahzadeh, S., Petersen, G.F., Alvisi, G., Forwood, J.K.(2026) Life Sci Alliance 9
- PubMed: 41672735
- DOI: https://doi.org/10.26508/lsa.202503543
- Primary Citation Related Structures:
9Y0R - PubMed Abstract:
Activating transcription factor 2 (ATF2) is a member of the AP-1 superfamily that regulates essential cellular processes through its activity as a nuclear transcription factor. Although ATF2 plays well-established roles in neurodevelopment, inflammation, and cancer, the mechanisms underlying its nuclear localisation remain poorly characterised. Here, we investigate the structural and functional basis of ATF2 nuclear import via the classical importin-α/β1 (IMPα/β1) pathway. Using quantitative in vitro binding assays, we demonstrate that ATF2 interacts with multiple IMPα paralogues. Fluorescence polarisation measurements reveal the highest binding affinity for IMPα1, with progressively weaker interactions observed for IMPα3, IMPα5, and IMPα7. Crystallographic analysis of ATF2 bound to IMPα1 identifies two basic clusters that are important for interaction: site 1 ( 353 EKRRK 357 ), which binds the major site of IMPα1, and site 2 ( 372 KRK 374 ), which binds the minor site. Mutation of key residues confirms the importance of both motifs, with site 1 contributing more substantially to binding. Quantitative confocal laser scanning microscopy analysis in HEK293A cells supports these findings, showing that mutation of both clusters is required to fully abolish ATF2 nuclear localisation. Inhibition of classical nuclear import using Bimax2 significantly reduces nuclear accumulation, whereas treatment with leptomycin B confirms a role of chromosomal region maintenance 1 (CRM1)-mediated nuclear export. Notably, ATF2 mutants incapable of nuclear import can localise to the nucleus when co-expressed with c-Jun, indicating that c-Jun can facilitate ATF2 nuclear import via heterodimerisation. Together, these results establish that ATF2 enters the nucleus through IMPα recognition of two basic clusters and highlight the redundancy and complexity of ATF2 nuclear trafficking mechanisms.
- School of Life and Environmental Sciences, University of Sydney, Sydney, Australia.
Organizational Affiliation: