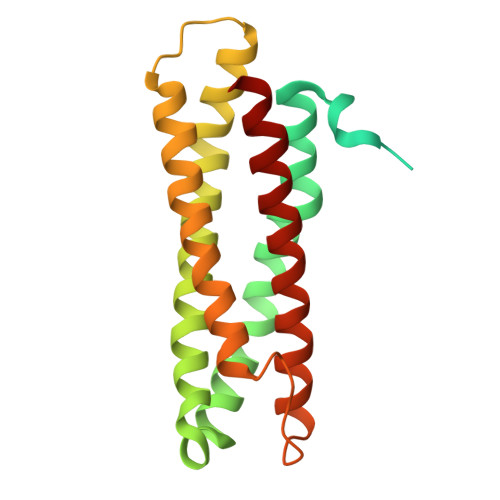
Structure and Nitrite Reductase Activity of the Di-iron Protein ScdA in Staphylococcus aureus.
Chen, H.Y., Tsai, R.F., Lu, Y.S., Cheng, Y.C., Fan-Chiang, H.Y., Wu, C.Y., Lo, F.C., Kuo, H.W., Yang, W.K., Liao, W.Y., Hu, N.J., Sue, S.C., Chiang, Y.W.(2025) J Am Chem Soc 147: 31558-31569
- PubMed: 40846682
- DOI: https://doi.org/10.1021/jacs.5c05573
- Primary Citation of Related Structures:
9J47, 9W8G - PubMed Abstract:
Pathogenic Staphylococcus aureus endures bursts of host-derived reactive nitrogen species, yet the molecular defenses that enable this resilience have remained unclear. We now show that the previously enigmatic di-iron enzyme ScdA functions as a nitrite reductase, converting nitrite to nitric oxide (NO), and we elucidate the structural elements that support this activity. Using an integrative toolkit─X-ray crystallography, solution NMR, AlphaFold modeling, and pulsed EPR/DEER─we solved the full-length homodimeric structure of ScdA and identified a robust di-iron center that forms stable iron-nitrosyl intermediates. Targeted mutagenesis reveals that redox-active cysteines and dimerization state tune catalytic output, whereas steady-state kinetics confirm efficient nitrite-to-NO turnover. In vivo, ScdA overexpression in Escherichia coli suppresses growth under nitrite-rich conditions, highlighting the cytotoxic potency of the NO it generates. By coupling structure to function, our work clarifies S. aureus strategies for managing nitrosylative stress and points to ScdA as a potential vulnerability in antibiotic-resistant pathogens.
- Department of Chemistry, National Tsing Hua University, Hsinchu 300-044, Taiwan.
Organizational Affiliation: