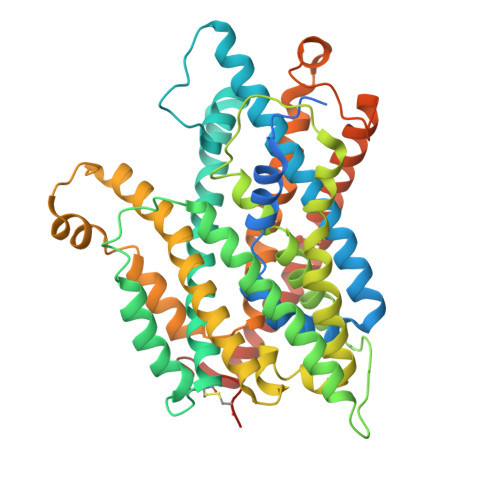
Structural basis of auxin recognition and transport by the plant influx carrier AUX1.
Chen, H., Fan, J., Chi, C., Zhao, J., Wu, D., Lei, X., Deng, X.W., Jiang, D.(2025) Mol Plant 18: 1284-1293
- PubMed: 40579814 Search on PubMed
- DOI: https://doi.org/10.1016/j.molp.2025.06.015
- Primary Citation Related Structures:
9UTD, 9UTE - PubMed Abstract:
Auxin regulates numerous aspects of plant growth and development, featuring polar auxin transport mediated by auxin efflux and influx carriers. AUX1 is the major auxin importer that actively takes up natural and synthetic auxins. However, the precise mechanisms underlying AUX1-mediated auxin recognition and transport remain elusive. Here, we present cryoelectron microscopy structures of Arabidopsis thaliana AUX1 in both apo and auxin-bound states, revealing the structural basis for auxin recognition. Structural analyses show that AUX1 assumes the LeuT-like fold in an inward-facing conformation and the auxin analog 2,4-D is recognized by polar residues located in the central cavity of AUX1. Furthermore, we identify a putative cation-binding site that contributes to stabilizing the inward-facing conformation. Interestingly, we reveal that His249 undergoes a substantial conformational shift, and its mutation completely abolishes transport activity, suggesting a crucial role for His249 in AUX1 gating. Collectively, this study provides a structural foundation for a deeper understanding of auxin influx by AUX1-like carriers.
- Peking University Institute of Advanced Agricultural Sciences, Shandong Laboratory of Advanced Agricultural Sciences at Weifang, Weifang, Shandong 261000, China; Beijing National Laboratory for Condensed Matter Physics, Laboratory of Soft Matter Physics, Institute of Physics, Chinese Academy of Sciences, Beijing 100190, China.
Organizational Affiliation: