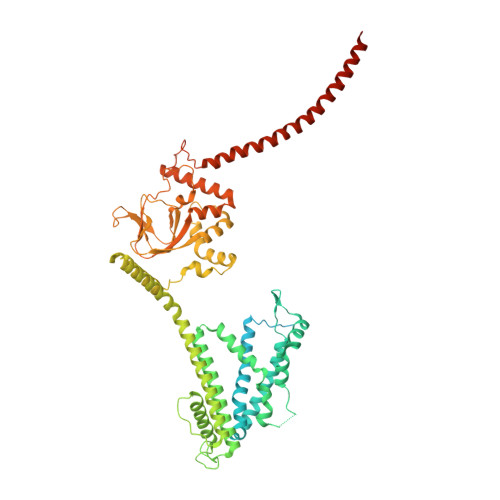
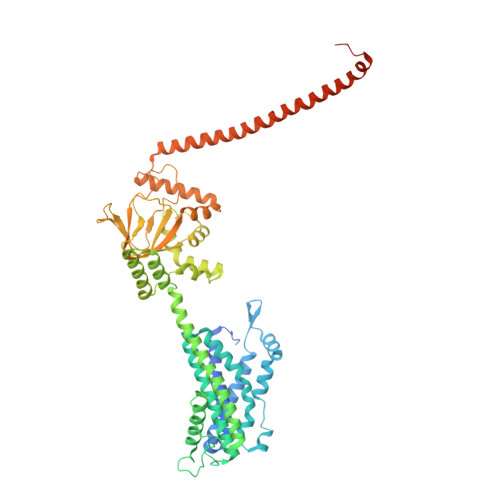
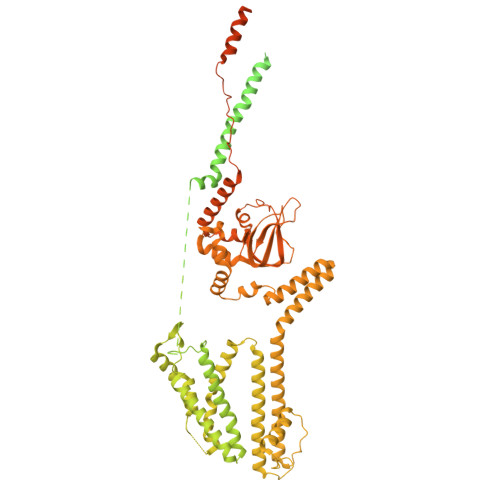
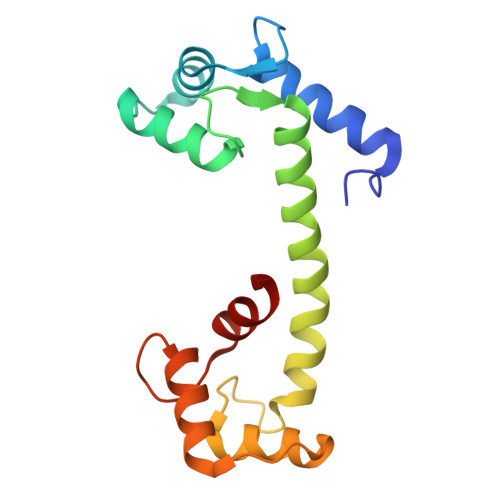
Structural mechanisms of assembly, gating, and calmodulin modulation of human olfactory CNG channel.
Xue, J., Gan, N., Zeng, W., Jiang, Y.(2025) Nat Commun 16: 9380-9380
- PubMed: 41131016 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-64436-5
- Primary Citation Related Structures:
9UPF, 9UPG - PubMed Abstract:
Mammalian cyclic nucleotide-gated (CNG) channels play crucial roles in visual and olfactory signal transduction. In olfactory sensory neurons, the native CNG channel functions as a heterotetramer consisting of CNGA2, CNGA4, and CNGB1b subunits and is activated by cAMP. Calmodulin (CaM) modulates the activity of the olfactory CNG channel, enabling rapid adaptation to odorants. Here we present cryo-EM structures of the native human olfactory CNGA2/A4/B1b channel in both CaM-bound closed and cAMP-bound open states, elucidating the molecular basis of the 2:1:1 subunit stoichiometry in channel assembly and the asymmetrical channel gating upon cAMP activation. Combining structural and functional analyses with AlphaFold prediction, we define two distinct CaM binding sites (CaM1 and CaM2) on the N- and C-terminal regions of CNGB1b, respectively, shedding light on the molecular mechanism of Ca 2+ /CaM-mediated rapid inhibition of the native olfactory CNG channel.
- Howard Hughes Medical Institute and Department of Physiology, University of Texas Southwestern Medical Center, Dallas, TX, USA. jxue@shsmu.edu.cn.
Organizational Affiliation: