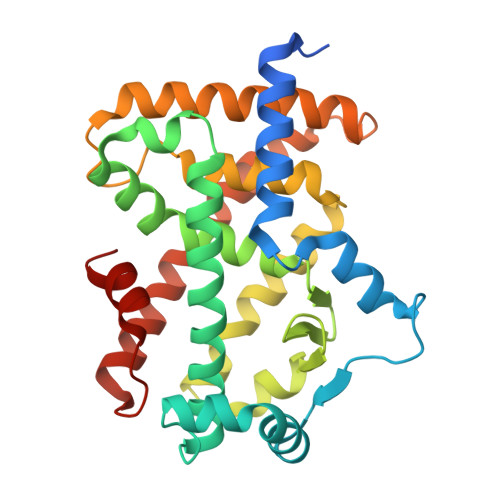
Bempedoic acid directly binds and activates PPAR alpha.
Papa, C., Rose, A., Martin, H.N.G., Useini, A., Geier, F., Liao, L., Rodriguez-Aguilera, J.R., Valina-Allo, P., Hoffmann, A., Tvardovskiy, A., Zulfqar, F., Zimmerman, A., Schicht, G., Ott, F., Korner, C., Engelmann, B., Rolle-Kampczyk, U., von Bergen, M., Meier, M., Bartke, T., Seehofer, D., Kloting, N., Matz-Soja, M., Damm, G., Boeckel, J.N., Buescher, J.M., Bluher, M., Laufs, U., Bondareva, O., Strater, N., Kunze, G., Heiker, J.T., Sheikh, B.N.(2026) Cell Metab
- PubMed: 41592562
- DOI: https://doi.org/10.1016/j.cmet.2025.12.018
- Primary Citation Related Structures:
9T5S - PubMed Abstract:
Bempedoic acid (BA) is a recently approved drug that lowers cholesterol and hepatic lipids, yet its mechanism of action remains incompletely understood. Here, we combine transcriptomic, biochemical, and structural approaches to show that BA directly binds to and activates peroxisome proliferator-activated receptor alpha (PPARα). BA treatment robustly induced PPARα signaling and fatty acid oxidation in primary hepatocytes and mouse liver. Through X-ray crystallography, we uncovered that BA binds to the ligand-binding domain of PPARα and stabilizes its active conformation. BA activated PPARα target genes independently of very-long-chain acyl-coenzyme A (CoA) synthetase (ACSVL1), the liver-enriched enzyme that converts BA to its bempedoyl-CoA form. Notably, BA-mediated induction of fatty acid oxidation required PPARα. Together, this work reveals direct PPARα activation as a key mechanism of BA action, providing a molecular basis for its lipid-lowering effects and suggesting broader therapeutic potential beyond the liver.
- Helmholtz Institute for Metabolic, Obesity and Vascular Research (HI-MAG), Helmholtz Center Munich, 04103 Leipzig, Germany.
Organizational Affiliation: