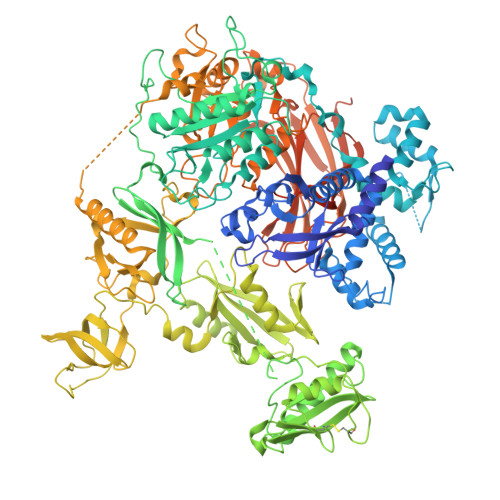
Characterisation of an allosteric site in PLC gamma enzymes and implications for development of their specific inhibitors.
Bunney, T.D., Nyvall, H.G., Macrae, C., Lalovic, D., Gregory, A., Le Huray, K.I.P., Harvey, N., Pinotsis, N., Kalli, A.C., Waudby, C.A., Burke, J.E., Katan, M.(2025) Biochem J 482: 1545-1564
- PubMed: 41002943
- DOI: https://doi.org/10.1042/BCJ20253358
- Primary Citation of Related Structures:
9QB7 - PubMed Abstract:
Phospholipase C gamma (PLCγ) enzymes are key components of intracellular signal transduction processes and are involved in disease development, including immune dysregulation, specific cancer types and neurodegeneration. Although recognised as important targets for intervention, validated pharmacological tools are lacking. Here, we demonstrate that inhibitory nucleotides bind directly to an allosteric site at the interface between the PLC-core and regulatory-array unique for PLCγ, underlying their specificity for the PLCγ family. This binding site overlaps with the PLCγ autoinhibitory interface, suggesting that the inhibitory impact of nucleotides involves stabilisation of autoinhibition. We have also analysed disease-linked variants of PLCγ1 and PLCγ2 to show that multiple mechanisms could underpin their gain-of-function phenotype. While the sensitivity of these variants to physiological nucleotide inhibition is reduced, we identified artificial nucleotide compounds that can inhibit such variants not only in vitro but also in cell-based assays. Therefore, our findings suggest a route for development of isozyme specific PLCγ inhibitors allowing further studies of their roles in health and disease.
- Institute of Structural and Molecular Biology, Division of Biosciences, University College London, London, WC1E 6BT, UK.
Organizational Affiliation: