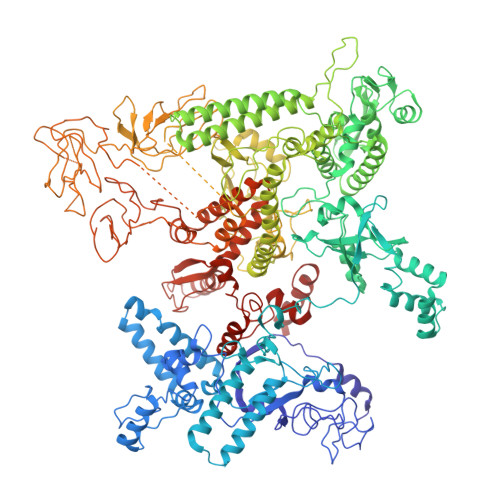
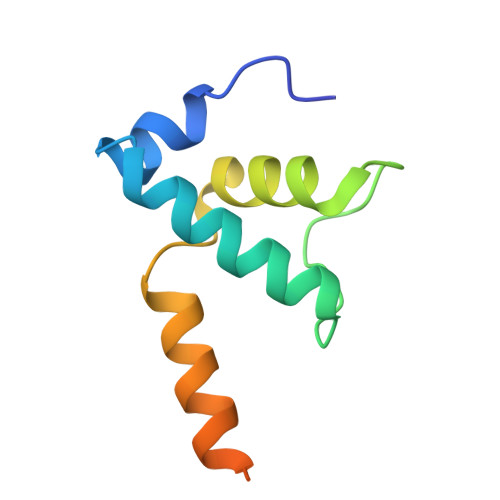
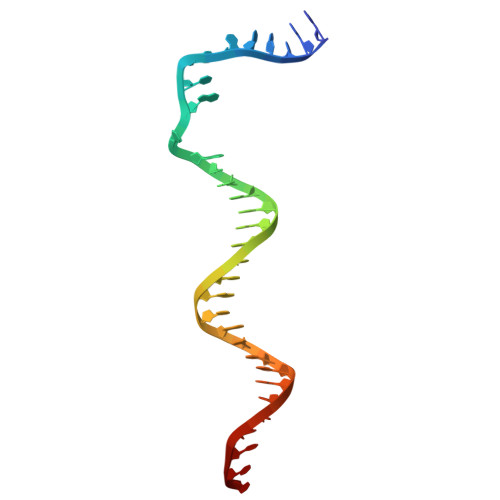
Subunit specialization in AAA+ proteins and substrate unfolding during transcription complex remodeling.
Gao, F., Ye, F., Buck, M., Zhang, X.(2025) Proc Natl Acad Sci U S A 122: e2425868122-e2425868122
- PubMed: 40273105 Search on PubMed
- DOI: https://doi.org/10.1073/pnas.2425868122
- Primary Citation Related Structures:
9Q90, 9Q91, 9Q92, 9Q93, 9Q94, 9Q95, 9Q96, 9Q97, 9Q98 - PubMed Abstract:
Bacterial RNA polymerase (RNAP) is a multisubunit enzyme that copies DNA into RNA in a process known as transcription. Bacteria use σ factors to recruit RNAP to promoter regions of genes that need to be transcribed, with 60% bacteria containing at least one specialized σ factor, σ 54 . σ 54 recruits RNAP to promoters of genes associated with stress responses and forms a stable closed complex that does not spontaneously isomerize to the open state where promoter DNA is melted out and competent for transcription. The σ 54 -mediated open complex formation requires specific AAA+ proteins ( A TPases A ssociated with diverse cellular A ctivities) known as bacterial enhancer-binding proteins (bEBPs). We have now obtained structures of new intermediate states of bEBP-bound complexes during transcription initiation, which elucidate the mechanism of DNA melting driven by ATPase activity of bEBPs and suggest a mechanistic model that couples the Adenosine triphosphate (ATP) hydrolysis cycle within the bEBP hexamer with σ 54 unfolding. Our data reveal that bEBP forms a nonplanar hexamer with the hydrolysis-ready subunit located at the furthest/highest point of the spiral hexamer relative to the RNAP. ATP hydrolysis induces conformational changes in bEBP that drives a vectoral transiting of the regulatory N terminus of σ 54 into the bEBP hexamer central pore causing the partial unfolding of σ 54 , while forming specific bEBP contacts with promoter DNA. Furthermore, our data suggest a mechanism of the bEBP AAA+ protein that is distinct from the hand-over-hand mechanism proposed for many other AAA+ proteins, highlighting the versatile mechanisms utilized by the large protein family.
- Section of structural and synthetic biology, Department of Infectious Disease, Imperial College London, London SW7 2AZ, United Kingdom.
Organizational Affiliation: