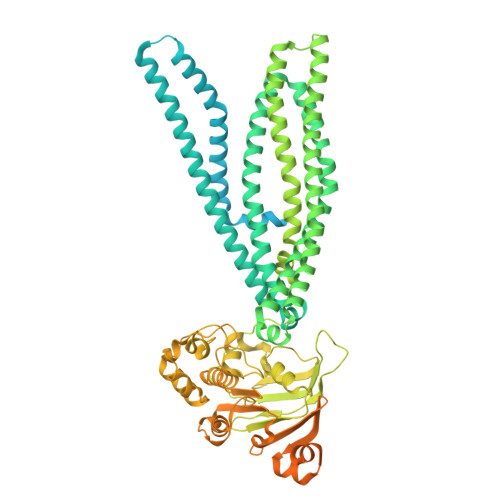
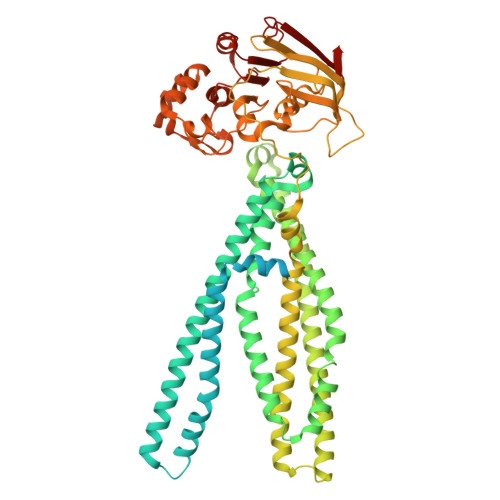
Structurally diverse viral inhibitors converge on a shared mechanism to stall the antigen transporter TAP.
Lee, J., Manon, V., Chen, J.(2025) Proc Natl Acad Sci U S A 122: e2516676122-e2516676122
- PubMed: 40956880 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2516676122
- Primary Citation Related Structures:
9O94, 9O9D, 9OCG, 9OCH, 9OCI, 9OCJ - PubMed Abstract:
In the host-pathogen arms race, herpesviruses and poxviruses encode proteins that sabotage the transporter associated with antigen processing (TAP), thereby suppressing MHC-I antigen presentation and enabling lifelong infection. Of the five known viral TAP inhibitors, only the herpes simplex virus (HSV) protein ICP47 has been structurally resolved. We now report cryoelectron microscopy structures of TAP in complex with the remaining four: BNLF2a (Epstein-Barr virus), hUS6 (human cytomegalovirus), bUL49.5 (bovine herpesvirus 1), and CPXV012 (cowpox virus), assembling a structural atlas of viral TAP evasion. Despite employing divergent sequences, folds, and conformational targets, these viral inhibitors converge on a common strategy: they stall TAP from the alternating access cycle, precluding peptide entry into the ER and shielding infected cells from cytotoxic T cell surveillance. These findings reveal striking functional convergence and provide a structural framework for rational antiviral design.
- Laboratory of Membrane Biophysics and Biology, The Rockefeller University, New York, NY 10065.
Organizational Affiliation: