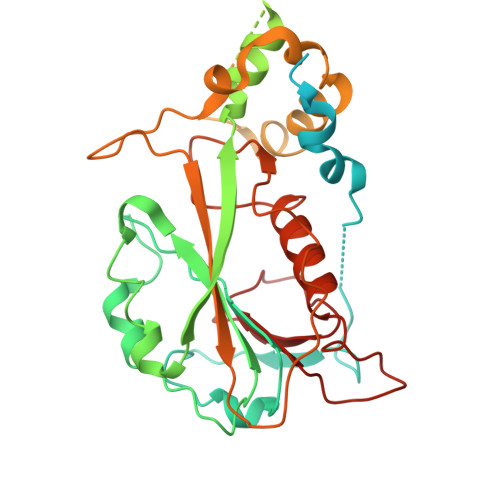
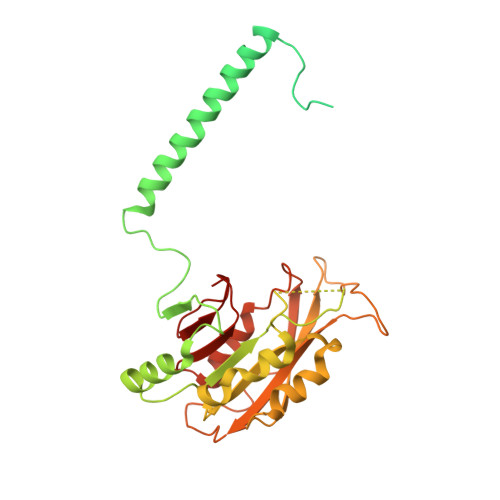
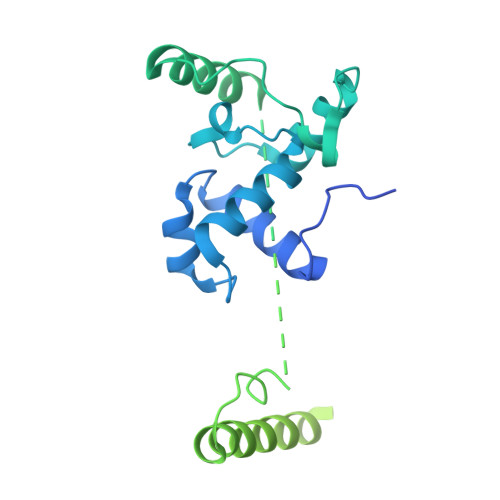
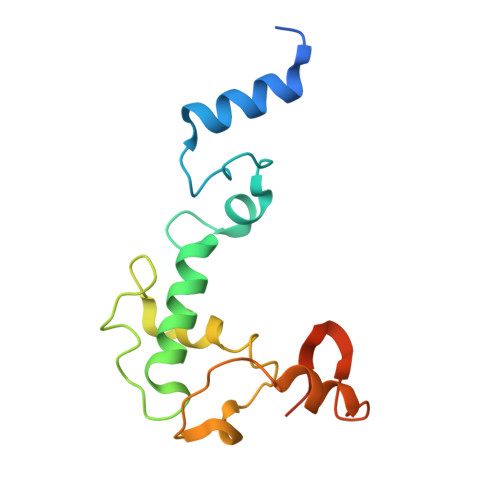
Structural insight into the substrate binding of the AMT complex via an inhibitor-trapped state.
Shao, Z., Yoon, S., Lu, J., Athavale, P., Liu, Y., Song, J.(2025) Protein Sci 34: e70265-e70265
- PubMed: 40815297
- DOI: https://doi.org/10.1002/pro.70265
- Primary Citation of Related Structures:
9O6K - PubMed Abstract:
N6-adenine (6mA) DNA methylation plays an important role in gene regulation and genome stability. The 6mA methylation in Tetrahymena thermophila is mainly mediated by the AMT complex, comprised of the AMT1, AMT7, AMTP1, and AMTP2 subunits. To date, how this complex assembles on the DNA substrate remains elusive. Here we report the structure of the AMT complex bound to the OCR protein from bacteriophage T7, mimicking the AMT-DNA encounter complex. The AMT1-AMT7 heterodimer approaches OCR from one side, while the AMTP1 N-terminal domain, assuming a homeodomain fold, binds to OCR from the other side, resulting in a saddle-shaped architecture reminiscent of what was observed for prokaryotic 6mA writers. Mutation of the AMT1, AMT7, and AMTP1 residues on the OCR-contact points led to impaired DNA methylation activity to various extents, supporting a role for these residues in DNA binding. Furthermore, structural comparison of the AMT1-AMT7 subunits with the evolutionarily related METTL3-METTL14 and AMT1-AMT6 complexes reveals sequence conservation and divergence in the region corresponding to the OCR-binding site, shedding light on the substrate binding of the latter two complexes. Together, this study supports a model in which the AMT complex undergoes a substrate binding-induced open-to-closed conformational transition, with implications in its substrate binding and processive 6mA methylation.
- Department of Biochemistry, University of California, Riverside, California, USA.
Organizational Affiliation: