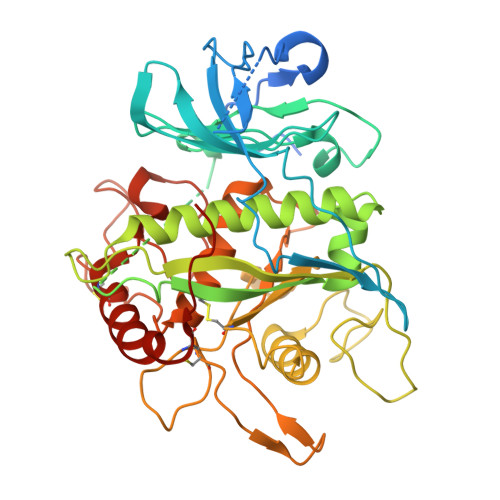
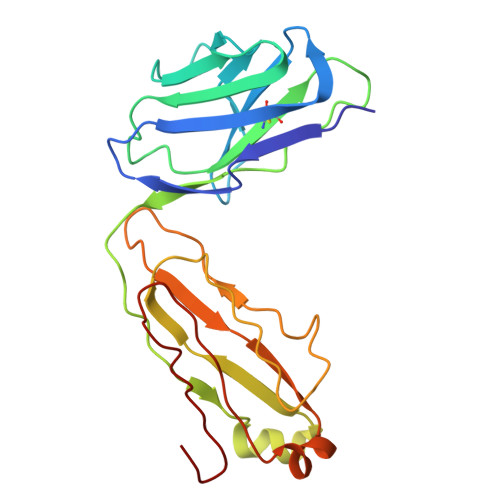
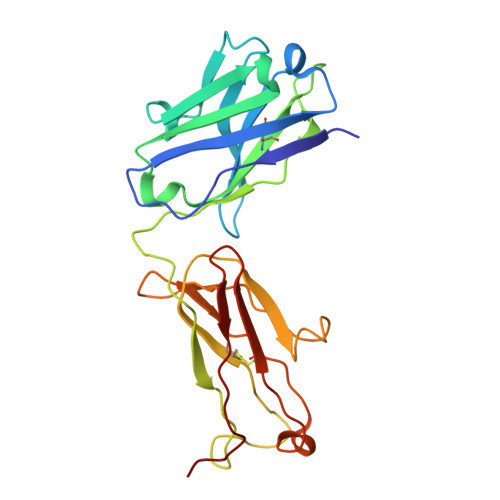
Structural insights into the activation and inhibition of the ADAM17-iRhom2 complex.
Maciag, J.J., Slone, C.E., Alnajjar, H.F., Rich, M.F., Guion, B., Ifergan, I., Blobel, C.P., Seegar, T.C.M.(2025) Proc Natl Acad Sci U S A 122: e2500732122-e2500732122
- PubMed: 40512800 Search on PubMed
- DOI: https://doi.org/10.1073/pnas.2500732122
- Primary Citation Related Structures:
9O54, 9O58 - PubMed Abstract:
The endopeptidase activity of ADAM (a disintegrin and metalloproteinase)-17, the primary processor of several EGFR ligands and tumor necrosis factor-alpha (TNF-α), is essential for proper embryonic development and immune regulation. Dysregulated ADAM17 activity is prevalent in a wide array of human diseases, including cancer, chronic inflammation, and SARS-CoV-2 viral progression. Initially translated as an inactive zymogen, ADAM17 maturation and enzymatic function are tightly regulated by its obligate binding partners, the inactive rhomboid proteins (iRhom) -1 and -2. Here, we present the cryo-EM structure of the ADAM17 zymogen bound to iRhom2. Our findings elucidate the interactions within the ADAM17-iRhom2 complex, the inhibitory mechanisms of the therapeutic MEDI3622 antibody and ADAM17 prodomain, and the previously unknown role of a membrane-proximal cytoplasmic reentry loop of iRhom2 involved in the mechanism of activation. Importantly, we perform cellular assays to validate our structural findings and provide further insights into the functional implications of these interactions, paving the way for developing therapeutic strategies targeting this biomedically critical enzyme complex.
- Department of Molecular and Cellular Biosciences, University of Cincinnati College of Medicine, Cincinnati, OH 45267.
Organizational Affiliation: