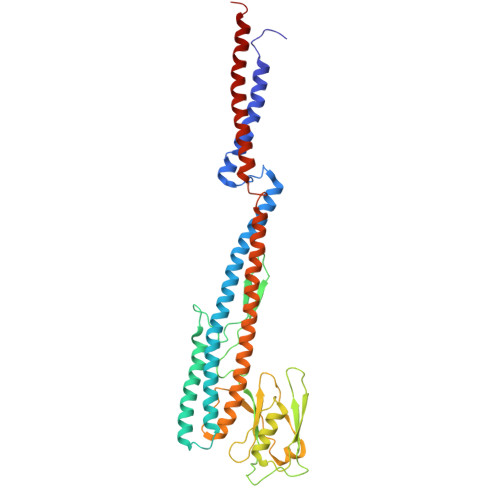
Structures of the sheathed flagellum reveal mechanisms of assembly and rotation in Vibrio cholerae.
Guo, W., Zhang, S., Park, J.H., Stanton, V., Asp, M., Herrera, H., Tai, J.B., Yue, J., Wang, J., Guo, J., Kumar, R., Botting, J.M., Wu, S., Yan, J., Klose, K.E., Yildiz, F.H., Liu, J.(2025) Nat Microbiol 10: 3305-3314
- PubMed: 41174224 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41564-025-02161-x
- Primary Citation Related Structures:
9N8A, 9N8B, 9N8G, 9N8H, 9N8M, 9P7R - PubMed Abstract:
Motility promotes the complex life cycle and infectious capabilities of Vibrio cholerae and is driven by rotation of a single polar flagellum. The flagellar filament comprises four flagellin proteins (FlaA-D) and is covered by a membranous sheath continuous with the outer membrane. Here we combine in situ cryo-electron microscopy single-particle analysis, fluorescence microscopy and molecular genetics to determine 2.92-3.43 Å structures of the sheathed flagellar filament from intact bacteria. Our data reveal the spatial arrangement of FlaA-D, showing that FlaA localizes at the cell pole and functions as a template for filament assembly involving multiple flagellins. Unlike unsheathed flagellar filaments, the sheathed filament from V. cholerae possesses a highly conserved core but a smooth, hydrophilic surface adjacent to the membranous sheath. A tiny conformational change at the single flagellin level results in a supercoiled filament and curved membranous sheath, supporting a model wherein the filament rotates separately from the sheath, enabling the distinct motility of V. cholerae.
- Department of Microbial Pathogenesis, Yale School of Medicine, New Haven, CT, USA. wangbiao.guo@yale.edu.
Organizational Affiliation: