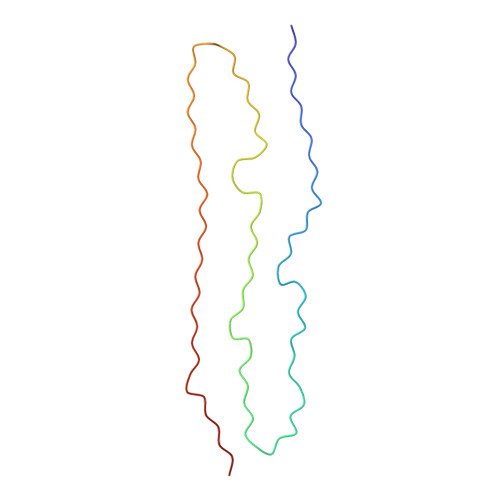Structures of Delta D421 Truncated Tau Fibrils.
El Mammeri, N., Duan, P., Hong, M.(2025) J Mol Biology 437: 169051-169051
- PubMed: 40021051 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2025.169051
- Primary Citation Related Structures:
9MR8 - PubMed Abstract:
The microtubule-associated protein tau aggregates into pathological β -sheet amyloid fibrils in Alzheimer's disease (AD) and other neurodegenerative diseases. In these aggregates, tau is chemically modified, including abnormal hyperphosphorylation and truncation. Truncation after D421 in the C-terminal domain occurs at early stages of AD. Here we investigate the structures of Δ D421-truncated 0N4R tau fibrils assembled in vitro in the absence of anionic cofactors. Using solid-state NMR spectroscopy and cryoelectron microscopy, we show that Δ D421-truncated 0N4R tau forms homogeneous fibrils whose rigid core adopts a three-layered β -sheet structure that spans R2, R3 and R4 repeats. This structure is essentially identical to that of full-length tau containing phospho-mimetic mutations at the PHF1 epitope in the C-terminal domain. In comparison, a Δ D421-truncated tau that additionally contains three phospho-mimetic mutations at the AT8 epitope in the proline-rich region forms a fibril core that includes the first half of the C-terminal domain, which is excluded from all known pathological tau fibril cores. These results indicate that the posttranslational modification code of tau contains redundancy: both charge modification and truncation of the C-terminal domain promote a three-layered β -sheet structure, which resembles pathological four-repeat tau structures in several tauopathies. In comparison, reducing the positive charges at the AT8 epitope in Δ D421-truncated tau promotes a fibril core that includes an immobilized C-terminal domain. The absence of this structure in tauopathy brains implies that Δ D421 truncation does not occur in conjunction with AT8 phosphorylation in diseased brains.
- Department of Chemistry, Massachusetts Institute of Technology, 170 Albany Street, Cambridge, MA 02139, United States.
Organizational Affiliation: