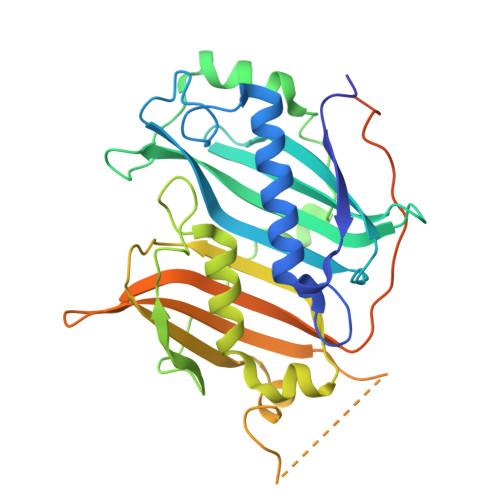
Structural Characterization of an Endogenous Algal Acyl-ACP Thioesterase.
Chen, J.A., Suo, Y., Mayfield, S.P., Burkart, M.D.(2025) Biochemistry 64: 3508-3514
- PubMed: 40768417 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.5c00088
- Primary Citation Related Structures:
9MQF - PubMed Abstract:
Fatty acids of specific chain lengths offer precursors for high-value renewable energy and fine chemicals industries. In plants and algae, the fatty acid chain length is determined by thioesterase-mediated hydrolysis of fatty acids from acyl carrier proteins through a hitherto unclear mechanism. Herein, a 2.50 Å resolution X-ray crystallography structure and an AlphaFold Multimer-generated model were used to identify active-site, substrate-binding, and protein-binding features contributing to catalysis. Coupled with mutational studies to determine impacts on product formation, we propose a catalytic mechanism involving water as a general base with surface residues specific to coordinating acyl carrier protein alignment. Binding tunnel restructuring altered substrate specificity of the thioesterase, and introduction of a non-native thioesterase with matching protein interface gave 95% hydrolysis of C12 fatty acids, offering new approaches for algae fatty acid biosynthetic design.
- Department of Chemistry and Biochemistry, University of California, San Diego, 9500 Gilman Drive, La Jolla, California 92093-0358, United States.
Organizational Affiliation: