Structure and function of a pair of non-competing monoclonal antibodies against Langya henipavirus attachment glycoprotein.
Li, W., Wu, S., Gui, Q., Liu, C., Zhou, B., Yan, H., Sun, Y., Fan, Q., Zhou, Y., Guo, H., Tang, S., Ge, X., Ju, B., Yan, R., Zhang, Z.(2025) Cell Rep 44: 116407-116407
- PubMed: 41071677 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2025.116407
- Primary Citation Related Structures:
9LEO - PubMed Abstract:
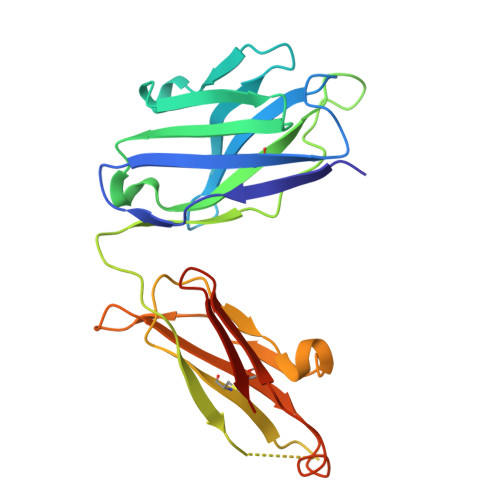
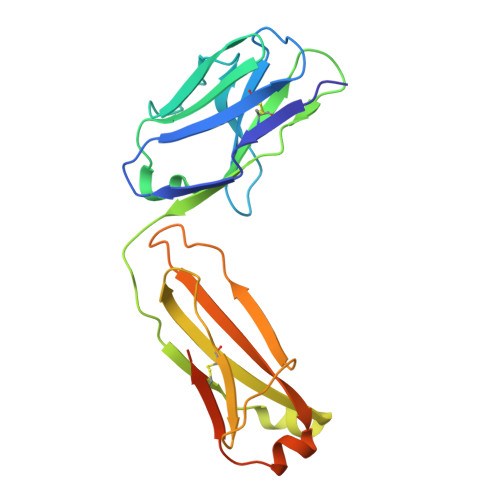
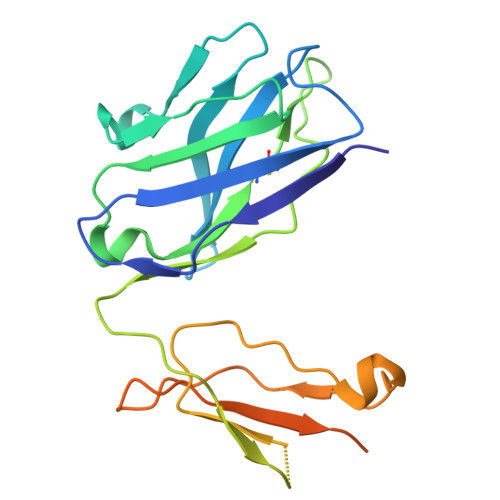
Langya henipavirus (LayV) is a zoonotic Parahenipavirus (Para-HNV) identified in recent years, discovered via surveillance of febrile patients with recent animal exposure in eastern China. The attachment glycoprotein (G) of HNV is critical for host cell entry and a key immune target. However, LayV-G exhibits notable antigenic differences from G of highly pathogenic bat-borne Hendra virus (HEV) and Nipah virus (NiV), implying vaccines or antibody therapies developed against HeV/NiV-G might be ineffective against LayV. Here, we immunize mice with LayV-G ectodomain and isolate a panel of LayV-G-targeting monoclonal antibodies (mAbs). We characterize two potent mAbs with pronounced crystallizable fragment (Fc)-mediated antiviral function and determine their cryo-electron microscopy (cryo-EM) structure binding to distinct epitopes of LayV-G head domain at a resolution of 2.92 Å, revealing antibody recognition mechanisms and potential conformational dynamics of LayV-G. Overall, our study defines two function-related epitopes of LayV-G, laying the foundation for therapeutic antibody development and vaccine design.
- Institute for Hepatology, National Clinical Research Center for Infectious Disease, Shenzhen Third People's Hospital, The Second Affiliated Hospital, School of Medicine, Southern University of Science and Technology, Shenzhen 518112, Guangdong, China.
Organizational Affiliation: