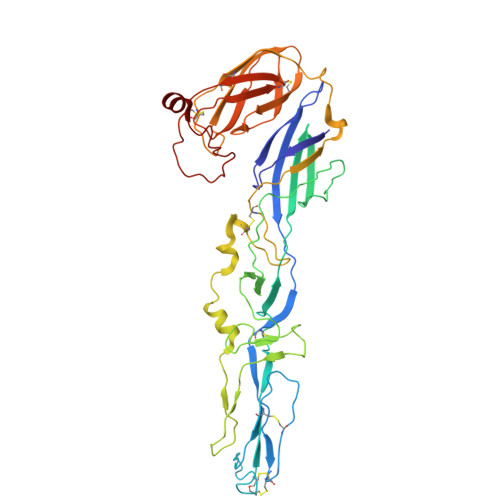
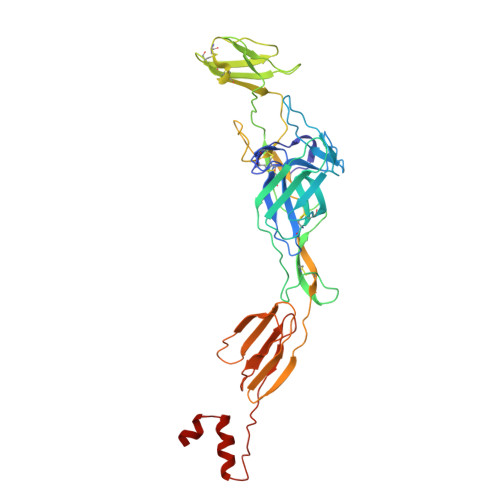
Structural basis for the recognition of two different types of receptors by Western equine encephalitis virus.
Ma, B., Cao, Z., Ding, W., Zhang, X., Xiang, Y., Cao, D.(2025) Cell Rep 44: 115724-115724
- PubMed: 40402741 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2025.115724
- Primary Citation Related Structures:
9L1N, 9L99, 9L9A - PubMed Abstract:
Western equine encephalitis virus (WEEV) enters cells via various receptors. Here, we report the cryoelectron microscopy (cryo-EM) structures of WEEV in complex with its receptors PCDH10 and very-low-density lipoprotein receptor (VLDLR). Structural analysis shows that PCDH10 binds in the cleft formed by adjacent E2-E1 heterodimers of WEEV through its EC1 ectodomain. Residues of viral envelope proteins involved in the interactions with PCDH10 EC1 are unique to WEEV. The strain-specific receptor VLDLR binds WEEV strain McMillan through two consecutive ecto-LDLR class A (LA) repeats. LA1-2, LA2-3, LA3-4, LA4-5, and LA5-6 of VLDLR all have detectable interactions with WEEV. Detailed structures of WEEV in complex with LA1-2 and LA2-3 show that the N-terminal LA repeat binds in the cleft and that the C-terminal LA repeat is attached to the E2 B domain. The acquisition of a single E2 mutation (V265F) allows WEEV strain 71V-1658, originally unable to bind VLDLR, to gain this receptor-binding ability. The binding of VLDLR to WEEV is in a mode different from those of other alphaviruses.
- Beijing Frontier Research Center for Biological Structure, Center for Infectious Disease Research, School of Basic Medical Sciences, Tsinghua University, Beijing 100084, P.R. China; SXMU-Tsinghua Collaborative Innovation Center for Frontier Medicine, Taiyuan 030001, P.R. China; Tsinghua-Peking Center for Life Sciences, Beijing 100084, P.R. China.
Organizational Affiliation: