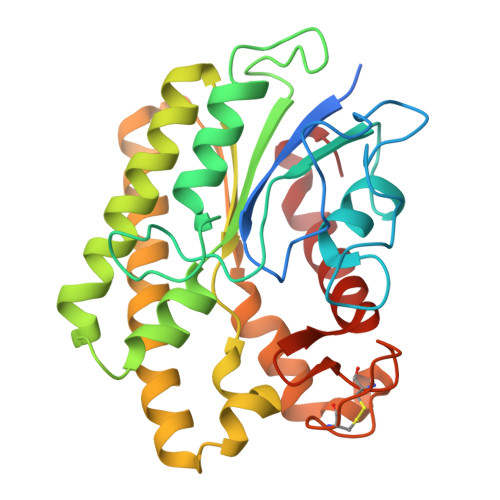
Structural and functional characterization of a novel GDSL-acetylesterase from Aspergillus niger reveals unique acylated choline specificity
Xing, S., He, L., Hu, G., Xie, W., Wang, L., Li, C., Tian, G., Wang, X., Yuan, Y., Gao, F., Liu, J.(2026) Chem Eng J 530: 173543