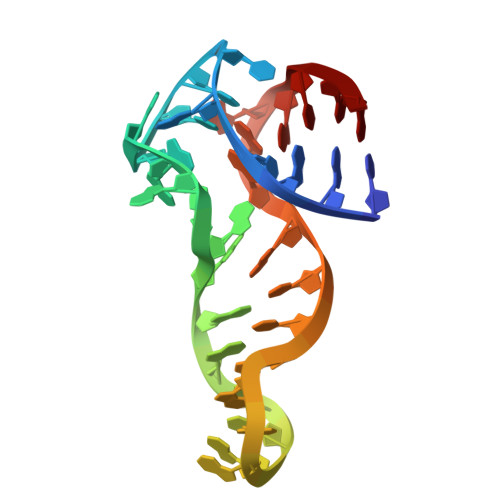
Structural basis and affinity improvement for an ATP-binding DNA aptamer.
Jiang, Y., Zhang, Y., Wan, L., Cui, C., Guo, P., Han, D., Tan, W.(2025) Proc Natl Acad Sci U S A 122: e2506491122-e2506491122
- PubMed: 40811466 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2506491122
- Primary Citation Related Structures:
9KTJ - PubMed Abstract:
DNA aptamers that bind small molecules with high affinity have revolutionized the fields of biosensing and bioimaging. Recently, a DNA aptamer named 1301b has been identified as the most potent DNA aptamer for the binding of adenosine triphosphate (ATP) with a dissociation constant ( K D ) of ~2.7 µM. However, the structural basis and recognition mechanism remain unclear, hindering further development of this DNA aptamer. In this study, we first design a shortened DNA aptamer namely 1301b_v1 that retains a good affinity for ATP and then determine the tertiary structure of 1:1 1301b_v1-ATP binding complex using solution NMR spectroscopy. The overall complex structure shows an "L" shape architecture with the binding pocket formed by two internal loops. The ATP intercalates into the binding pocket through forming hydrogen bond with G14 and stacking with T8·A28 and G9. We also reveal an adaptive binding mechanism where the DNA aptamer switches from a semifolded state to a stable tertiary structure upon ATP binding. Based on the structure-function relationship, we introduce 2'-O-methyl modification to residues in the central junction and obtain a DNA aptamer named 9/10/16 OMe with a K D of ~0.7 µM for the binding of ATP. These results underscore the ability of DNA molecules to form intricate three-dimensional folds with sophisticated functionality, opening up avenues for designing novel DNA-based molecular tools.
- Molecular Science and Biomedicine Laboratory, State Key Laboratory of Chemo and Biosensing, College of Chemistry and Chemical Engineering, College of Biology, Aptamer Engineering Center of Hunan Province, Hunan University, Changsha, Hunan 410082, China.
Organizational Affiliation: