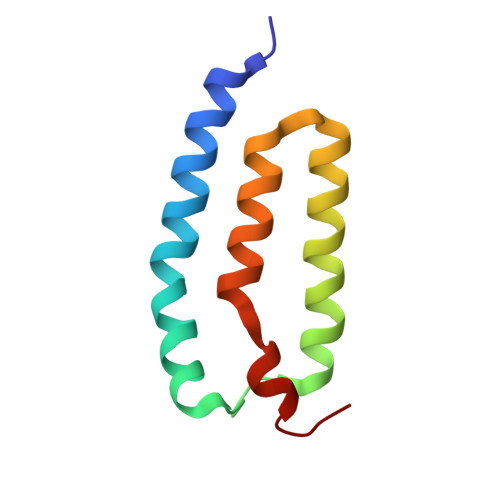
Dynamic structures of a membrane transporter in native cellular membranes.
Xie, H., Gan, Y., Zhao, W., Duan, M., Shen, Y., Tan, H., Zhang, Y., Tong, Q., Zhao, Y., Yang, J.(2025) Sci Adv 11: eadv4583-eadv4583
- PubMed: 41223279
- DOI: https://doi.org/10.1126/sciadv.adv4583
- Primary Citation Related Structures:
9KAX, 9KBA - PubMed Abstract:
Substrate transport through membrane transporters is a dynamic process that involves transitions between different functional conformations, which can be disrupted by non-native membrane mimetics. Capturing these conformations and their transitions within native cellular membranes presents a notable challenge. Herein, we used in situ solid-state nuclear magnetic resonance (NMR) to resolve the 1.5-Å outward-open and 2.5-Å occluded structures of Bj SemiSWEET within its native cellular membranes. Our findings reveal that these two conformations exchange within transmembrane helix TM1 and Loop L2-3 on a millisecond to second timescale, with the exchange rate corresponding to the sucrose transport rate. Molecular dynamics simulations further confirmed that these conformations represent functional states during sucrose transport. In contrast, we observed different conformational dynamics of Bj SemiSWEET in DMPC/DMPG synthetic bilayers compared to cellular membranes. This study highlights the potential of in situ solid-state NMR to provide previously unknown dynamic structural insights into cellular molecular processes, representing a substantial advancement in dynamic cellular structural biology.
- National Center for Magnetic Resonance in Wuhan, Key Laboratory of Magnetic Resonance in Biological Systems, State Key Laboratory of Magnetic Resonance and Atomic and Molecular Physics, Wuhan Institute of Physics and Mathematics, Innovation Academy for Precision Measurement Science and Technology, Chinese Academy of Sciences, Wuhan 430071, P. R. China.
Organizational Affiliation: