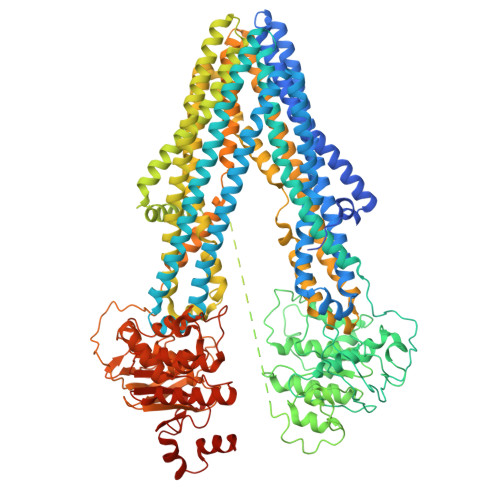
Structures and mechanisms of the ABC transporter ABCB1 from Arabidopsis thaliana.
Chen, Q., Zhu, L., Zhang, S., Qiao, S., Ding, Z.J., Zheng, S.J., Guo, J., Su, N.(2025) Structure 33: 903-915.e5
- PubMed: 40101709 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2025.02.008
- Primary Citation Related Structures:
9JUJ, 9JUK, 9JUL, 9JUM, 9JUN, 9JUO, 9JUP - PubMed Abstract:
The Arabidopsis thaliana auxin transporter ABCB1 plays a fundamental role in the regulation of plant growth and development. While its homolog ABCB19 was previously shown to transport brassinosteroids (BR), another class of essential hormones, the ability of ABCB1 to mediate BR transport has remained unexplored. In this study we show that ABCB1 also transports brassinosteroids with an in vitro brassinolide (BL) transport assay. Using single-particle cryo-electron microscopy, we determined ABCB1 structures in multiple inward-facing conformations in the apo state, ANP-bound state, BL-bound state, and the both BL- and ANP-bound state. BL binds to the large cavity of two transmembrane domains, inducing a slight conformational change. Additionally, we obtained the structure of ABCB1 in an outward-facing conformation. By comparing these different conformations, we elucidated the possible mechanism of hormone transport by ABCB1. These high-resolution structures help us to understand the structural basis for hormone recognition and transport mechanisms of ABCB1.
- Center for Membrane Receptors and Brain Medicine, the Fourth Affiliated Hospital of School of Medicine, and International School of Medicine, International Institutes of Medicine, Zhejiang University, Yiwu, Zhejiang 322000, China; Department of Biophysics and Department of Neurology of the Fourth Affiliated Hospital, Zhejiang University School of Medicine, Hangzhou, Zhejiang 310058, China; Nanhu Brain-computer Interface Institute, Hangzhou, Zhejiang 311100, China.
Organizational Affiliation: