Cryo-EM structure of MAS1-Gi complex
Wong, T.S., Xiong, T.T., Zeng, Z.C.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Guanine nucleotide-binding protein G(i) subunit alpha-1 | 340 | Homo sapiens | Mutation(s): 0 Gene Names: GNAI1 EC: 3.6.5 | 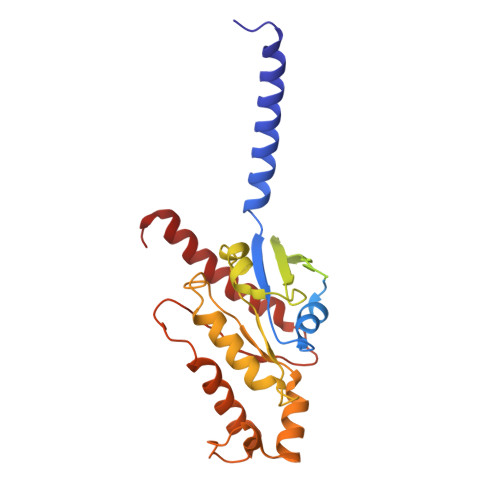 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P63096 GTEx: ENSG00000127955 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P63096 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 | 338 | Homo sapiens | Mutation(s): 0 Gene Names: GNB1 |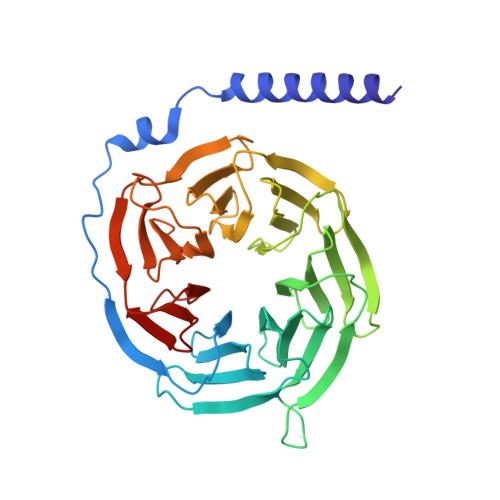  | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P62873 GTEx: ENSG00000078369 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P62873 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 | C [auth G] | 53 | Homo sapiens | Mutation(s): 0 Gene Names: GNG2 | 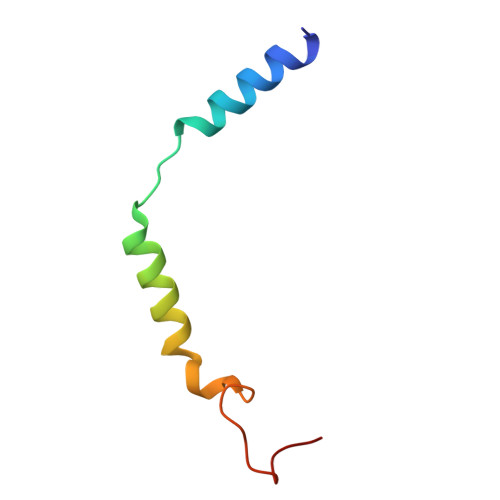 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P59768 GTEx: ENSG00000186469 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P59768 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Proto-oncogene Mas | D [auth R] | 253 | Homo sapiens | Mutation(s): 0 Gene Names: MAS1, MAS | 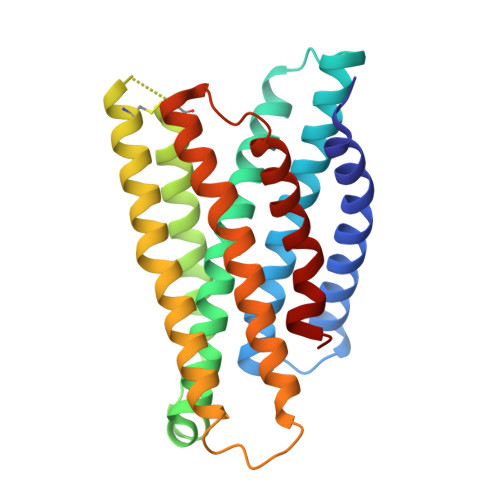 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P04201 GTEx: ENSG00000130368 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P04201 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| scFv16 | E [auth S] | 259 | Homo sapiens | Mutation(s): 0 | 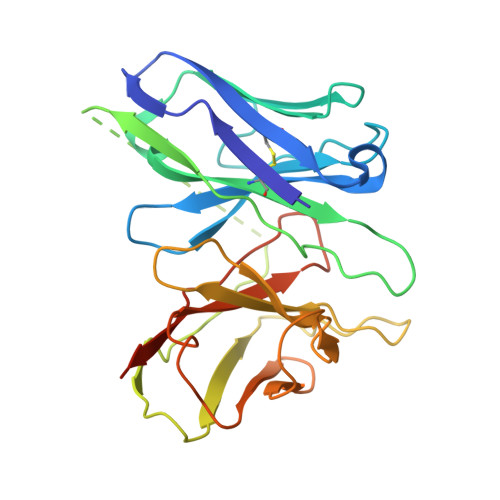 |
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| A1ECD (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | F [auth R] | 1-ethyl-3-[3-[4-[(5-methanoyl-4-methoxy-2-phenyl-imidazol-1-yl)methyl]phenyl]-5-(2-methylpropyl)thiophen-2-yl]sulfonyl-urea C29 H32 N4 O5 S2 QTOZBSNPDCWHPV-UHFFFAOYSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Not funded | -- |