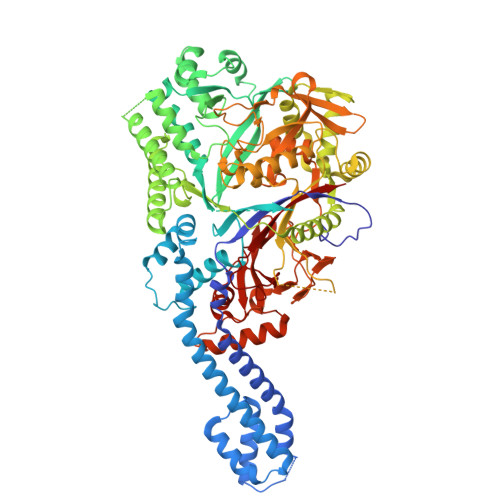
Structural Analysis of GodF, an O-glutamylation Enzyme Involved in Goadsporin Biosynthesis.
Shimizu-Ibuka, A., Kato, Y., Asamizu, S., Onaka, H.(2026) J Mol Biology 438: 169619-169619
- PubMed: 41478604
- DOI: https://doi.org/10.1016/j.jmb.2025.169619
- Primary Citation of Related Structures:
9JHQ - PubMed Abstract:
Goadsporin is one of linear azole-containing peptides (LAPs) that form a subgroup within ribosomally synthesized and post-translationally modified peptides (RiPPs). It contains two dehydroalanine residues formed through the action of two enzymes, GodF and GodG, in a two-step process involving serine O-glutamylation followed by elimination. Here, we report the X-ray crystal structure of GodF, which catalyzes the tRNA-dependent glutamylation of target serine residues, resolved at a 2.34-Å resolution. Although GodF exhibits low homology at the primary sequence level, its overall structure closely resembles that of TbtB, a tRNA Glu -dependent enzyme involved in thiopeptide biosynthesis, as well as the O-glutamylation domains of NisB and MibB, which serve as dehydroalanine synthases in lanthipeptide biosynthesis. The residues and structural elements forming the active site are well-aligned among these enzymes, while regions outside the active site are poorly conserved. Like TbtB, GodF features a coiled-coil subdomain at its N-terminus, and AlphaFold3 predicts this region plays a key role in recognizing the substrate tRNA Glu . GodF also contains a typical RiPP recognition element (RRE) motif; however, the spatial arrangement of the secondary structural elements comprising this motif differs notably from those in other O-glutamylating enzymes. These structural characteristics of GodF highlight the diversity of substrate-binding pockets among RiPP-modifying enzymes, reflecting the variability in their substrate peptides and the necessity to accommodate distinct conformational and physicochemical properties.
- Graduate School of Science, Kanagawa University, 3-27-1, Rokkakubashi, Kanagawa-ku, Yokohama 221-8686, Japan. Electronic address: ibuka@kanagawa-u.ac.jp.
Organizational Affiliation: