Multiscale biased chemical space remodeling for developing APLNR agonists with anti-HFpEF efficacy.
Sun, Q., Tian, X., Tan, L., Deng, Y., Liu, S., Xiong, Y., Feng, Y., Wang, Y., Zhang, L., Zhu, J., Xiao, H., Shao, Z., Guo, Y., Yan, W., Li, T., Ouyang, L.(2025) Proc Natl Acad Sci U S A 122: e2423432122-e2423432122
- PubMed: 40314976 Search on PubMed
- DOI: https://doi.org/10.1073/pnas.2423432122
- Primary Citation Related Structures:
9JH3 - PubMed Abstract:
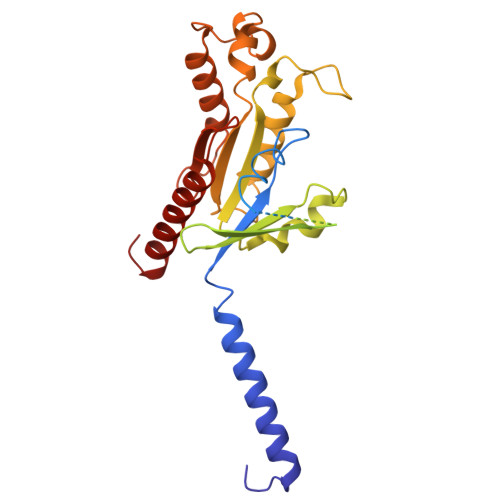
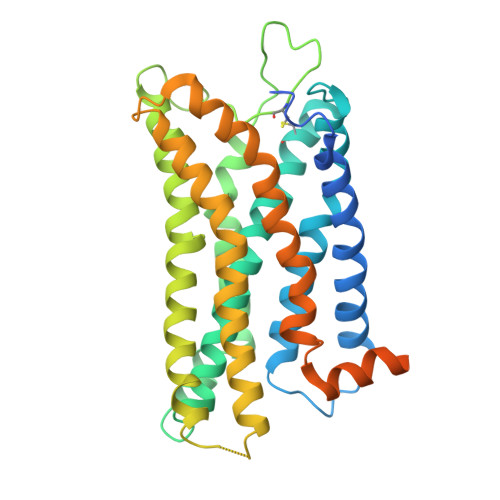
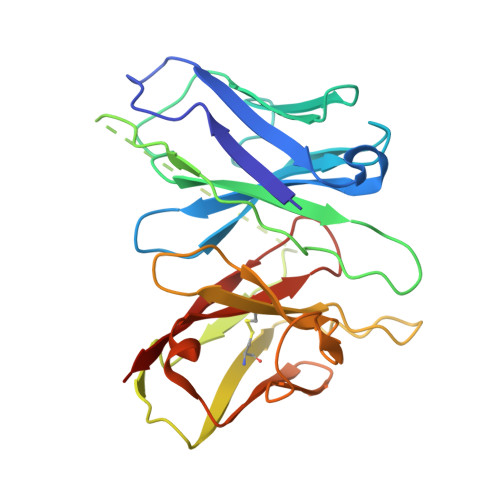
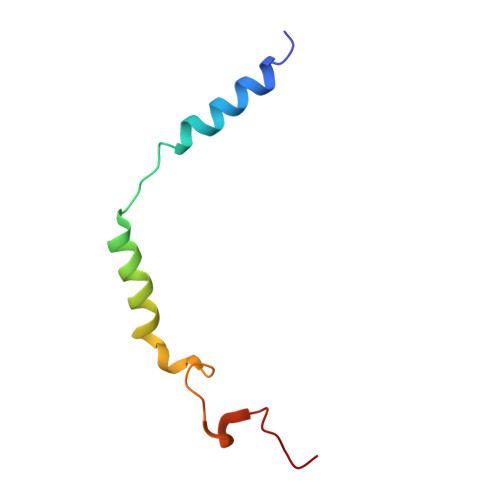
Heart failure with preserved ejection fraction (HFpEF) represents a significant global health burden, yet effective pharmacotherapies remain elusive. The angiotensin-like 1 receptor, also known as the apelin receptor (APLNR), is a promising target for treating HFpEF due to its role in modulating cardiovascular function. Despite the cardioprotective effects of endogenous ligand, apelin, achieving G-protein-biased agonism for therapeutic benefit poses a significant challenge. In this study, we unravel the biased signal transduction pathway mediated by a reported partial G i -protein-biased APLNR agonist CMF-019 and developed a biased chemical space remodeling approach to identify exclusive G-protein-biased agonists targeting APLNR. These agonists exhibited enhanced Gi-protein-biased function and protective effects in both in vitro and in vivo. Our findings not only enhance comprehension of APLNR-biased agonism but also establish drug design strategies for modifying and reshaping biased chemical landscapes in other G-protein-coupled receptors (GPCRs).
- Department of Cardiovascular Surgery, Cardiovascular Surgery Research Laboratory, State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu 610041, China.
Organizational Affiliation: