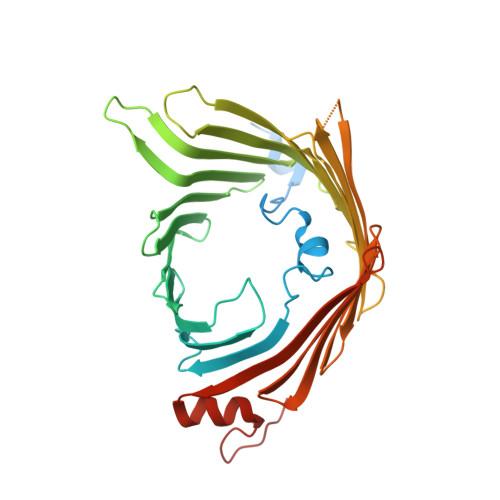
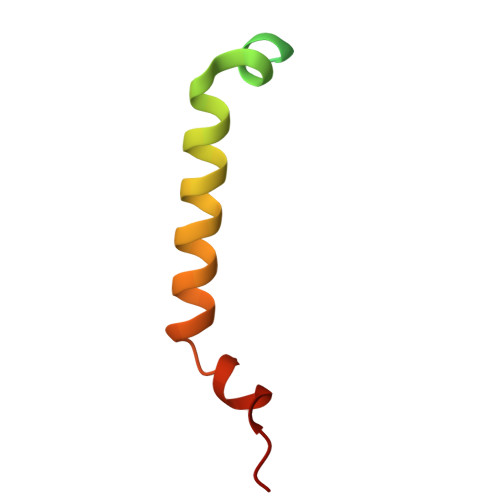
Dynamic TOM-TIM23 supercomplex directs mitochondrial protein translocation and sorting.
Yang, Y., Wang, S., Wang, G., Lian, Y., Xue, L., Jiang, W., Guo, Q., Song, C., Li, L.(2025) Nat Struct Mol Biol 32: 2231-2241
- PubMed: 40877479 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-025-01662-x
- Primary Citation Related Structures:
9J99, 9J9B - PubMed Abstract:
The mitochondrial translocase of the outer membrane (TOM) and translocase of the inner membrane 23 (TIM23) complexes are coupled to control protein import across the outer and inner membranes, respectively. However, the mechanisms of protein recognition and sorting in the TOM-TIM23 pathway remain unclear. Here we report cryo-electron microscopy structures of a translocating polypeptide substrate captured in the active TOM-TIM23 supercomplex from Saccharomyces cerevisiae. In the TOM complex, the polypeptide substrate adopts multiple conformations stabilized by hydrophilic residues from distinct regions of the Tom40 channel. In the TIM23 complex, the Tim17 and Mgr2 subunits create the translocation pathway, with a central restriction formed by four highly conserved hydrophobic residues. The substrate primarily interacts with hydrophobic residues along the Tim17-Mgr2 pathway. Substrate hydrophobicity modulates the association of Mgr2 with Tim17, enabling dynamic regulation of protein sorting toward either the matrix or membrane. These findings reveal a sophisticated translocation mechanism of the TOM-TIM23 supercomplex that ensures the efficient import of diverse mitochondrial proteins.
- State Key Laboratory of Membrane Biology, School of Life Sciences, Peking University, Beijing, China.
Organizational Affiliation: