Response to: The mechanism for GTP-mediated RNA capping by the SARS-CoV-2 NiRAN domain remains unresolved.
Huang, Y., Tan, L., Liu, Y., Zhao, H., Wang, J., Ge, J., Ye, S., Liu, Z., Lan, W., Huang, B., Zhang, H., Gao, Y., Yan, L., Rao, Z., Lou, Z.(2025) Cell 188: 4462-4469.e9
- PubMed: 40570837 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2025.05.045
- Primary Citation Related Structures:
9IKZ - PubMed Abstract:
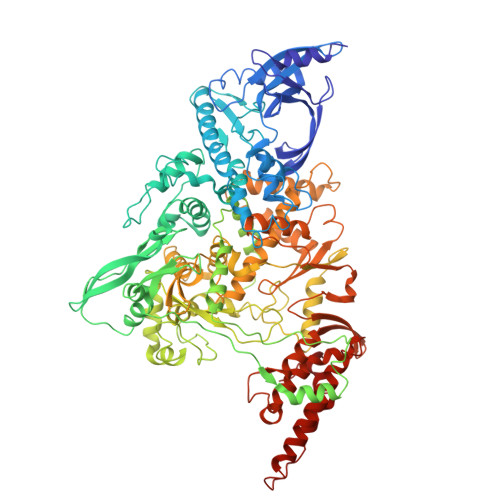
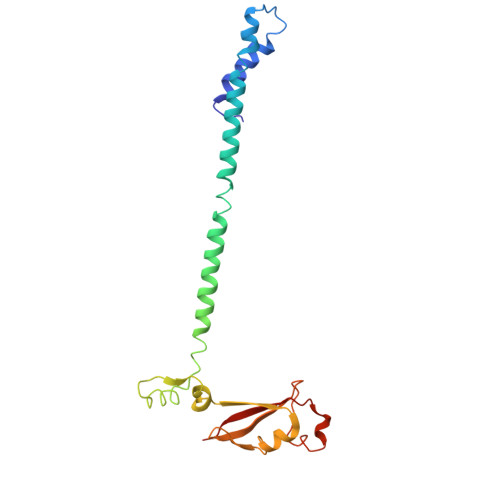
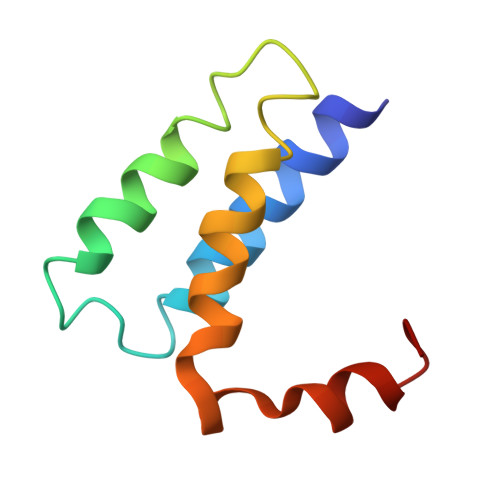
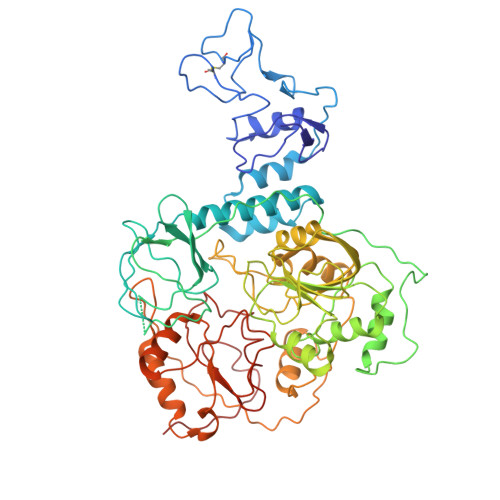
The SARS-CoV-2 polymerase NiRAN domain initiates RNA capping. Previous results showed that both GTP and GDP can be utilized by NiRAN to yield GpppA together with RNAylated nsp9 (RNA-nsp9); however, the G-pocket substrate selection and the working mechanism of NiRAN remain unclear. Small et al. questioned the binding of the non-hydrolyzable GTP analog GMPPNP in the G-pocket of the RTC:RNA-nsp9:GMPPNP structure (PDB: 8GWE) and proposed that the GTP-mediated RNA-capping mechanism remains unresolved. Here, we show the optimized density derived from the original data to support the modeling of GMPPNP, and we reveal why the alternative data processing method failed to obtain density results. We provide additional biochemical and structural evidence by using GTP, GDP, GMPPNP, and GDP⋅BeF 3 - as probes to clarify the GTP-mediated RNA-capping mechanism and reconcile the two currently known models by using GTP and GDP as substrates. This Matters Arising Response addresses the Small et al. (2025) Matters Arising paper, published concurrently in Cell.
- State Key Laboratory of Medicinal Chemical Biology, College of Life Sciences and College of Pharmacy, Nankai University, Tianjin, China.
Organizational Affiliation: