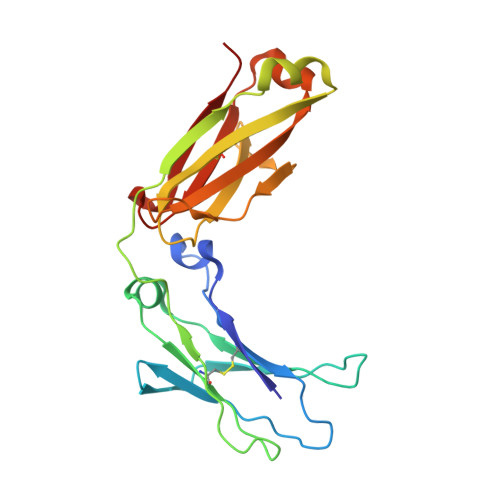
Experimental and Computational Insights into the Structural Dynamics of the Fc Fragment of IgG1 Subtype from Biosimilar VEGF-Trap
Destan, E., Turkut, E., Aldeniz, A., Kang, J., Tosha, T., Yabashi, M., Yilmaz, B., Timucin, C., Matsuura, H., Kawano, Y., Cinkaya, I., Demirci, H.(2025) Small Struct