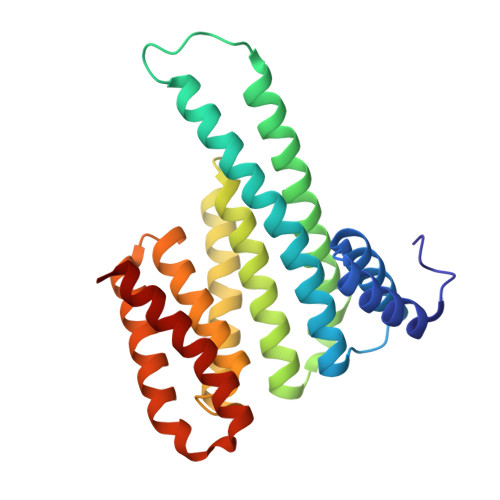
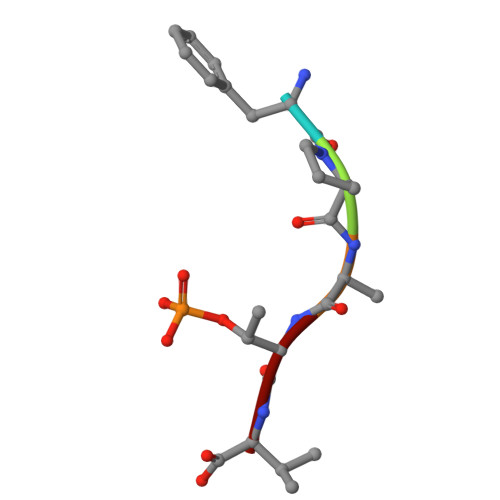
Scaffold-hopping for molecular glues targeting the 14-3-3/ER alpha complex.
Konstantinidou, M., Zingiridis, M., Pennings, M.A.M., Fragkiadakis, M., Virta, J.M., Revalde, J.L., Visser, E.J., Ottmann, C., Brunsveld, L., Neochoritis, C.G., Arkin, M.R.(2025) Nat Commun 16: 6467-6467
- PubMed: 40659654 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-61176-4
- Primary Citation Related Structures:
9I6S, 9I6T, 9I6U, 9I6V, 9I6W, 9I6X, 9I6Y, 9I6Z, 9I70, 9I71, 9I72, 9I73, 9I74, 9I75 - PubMed Abstract:
Molecular glues, small molecules that bind cooperatively at a protein-protein interface, have emerged as powerful modalities for the modulation of protein-protein interactions (PPIs) and "undruggable" targets. The systematic identification of new chemical matter with a molecular glue mechanism of action remains a significant challenge in drug discovery. Here, we present a scaffold hopping approach, using as a starting point our previously developed molecular glues for the native 14-3-3/estrogen receptor alpha (ERα) complex. The novel, computationally designed scaffold is based on the Groebke-Blackburn-Bienaymé multi-component reaction (MCR), leading to drug-like analogs with multiple points of variation, thus enabling the rapid derivatization and optimization of the scaffold. Structure-activity relationships (SAR) are developed using orthogonal biophysical assays, such as intact mass spectrometry, TR-FRET and SPR. Rational structure-guided optimization is facilitated by multiple crystal structures of ternary complexes with the glues, 14-3-3 and phospho-peptides mimicking the highly disordered C-terminus of ERα. Cellular stabilization of 14-3-3/ERα for the most potent analogs is confirmed using a NanoBRET assay with full-length proteins in live cells. Our approach highlights the potential of MCR chemistry, combined with scaffold hopping, to drive the development and optimization of unprecedented molecular glue scaffolds.
- Department of Pharmaceutical Chemistry and Small Molecule Discovery Centre (SMDC) University of California San Francisco (UCSF), San Francisco, CA, USA. markella.konstantinidou@ucsf.edu.
Organizational Affiliation: