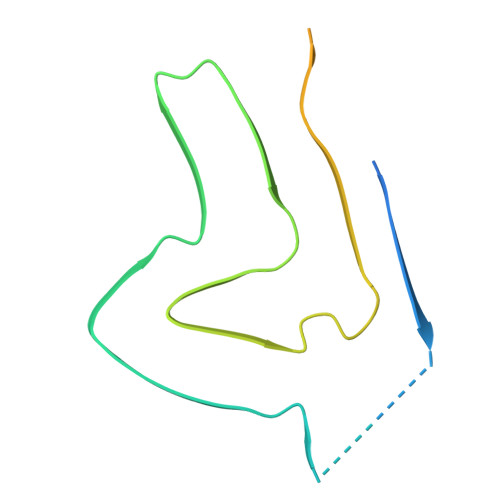
Structural insights into binding of novel PET tracer MODAG-005 to lipidic alpha-synuclein fibrils
Kim, M., Matthes, D., Frieg, B., Leonov, A., Ryazanov, S., Bleher, D., Grotegerd, A.-K., Dienemann, C., Giese, A., Schroeder, G.F., Becker, S., Herfert, K., de Groot, B.L., Andreas, L.B., Griesinger, C.To be published.